2025
|
Hiensch, Fleur; Dupré, Loïc; Salzer, Elisabeth Immunoactinopathies revisited: understanding clinical manifestations and biological pathways Journal Article In: Blood, vol. 145, no. 23, pp. 2709–2732, 2025, ISSN: 1528-0020. @article{hiensch_immunoactinopathies_2025,
title = {Immunoactinopathies revisited: understanding clinical manifestations and biological pathways},
author = {Fleur Hiensch and Loïc Dupré and Elisabeth Salzer},
doi = {10.1182/blood.2024026763},
issn = {1528-0020},
year = {2025},
date = {2025-06-01},
urldate = {2025-06-01},
journal = {Blood},
volume = {145},
number = {23},
pages = {2709–2732},
abstract = {Immune cell functionality is highly dependent on the actin cytoskeleton. The actin cytoskeleton is regulated by a complex molecular machinery that involves multiple genes. Mutations in these genes can cause inborn errors of immunity, also termed immunoactinopathies, of which Wiskott-Aldrich syndrome is the best-characterized entity. Currently, mutations in 23 genes can be considered causative of immunoactinopathies. Immunoactinopathies are rare disease entities with complex combinations of clinical manifestations, including immunodeficiency, immune dysregulation, malignancies, atopy, thrombocytopenia and bleeding, skin involvement, or congenital defects. Prompt diagnosis is crucial, because hematopoietic stem cell transplantation in an early phase can offer cure and prevent further complications. This review provides a detailed summary of the clinical experience with immunoactinopathies so far, elaborates on the most distinguishing features of immunoactinopathies by providing a clinical categorization, and links this information to the underlying biological pathways. This information may be of help to clinicians in the diagnosis of patients and to eventually improve patient care.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
Immune cell functionality is highly dependent on the actin cytoskeleton. The actin cytoskeleton is regulated by a complex molecular machinery that involves multiple genes. Mutations in these genes can cause inborn errors of immunity, also termed immunoactinopathies, of which Wiskott-Aldrich syndrome is the best-characterized entity. Currently, mutations in 23 genes can be considered causative of immunoactinopathies. Immunoactinopathies are rare disease entities with complex combinations of clinical manifestations, including immunodeficiency, immune dysregulation, malignancies, atopy, thrombocytopenia and bleeding, skin involvement, or congenital defects. Prompt diagnosis is crucial, because hematopoietic stem cell transplantation in an early phase can offer cure and prevent further complications. This review provides a detailed summary of the clinical experience with immunoactinopathies so far, elaborates on the most distinguishing features of immunoactinopathies by providing a clinical categorization, and links this information to the underlying biological pathways. This information may be of help to clinicians in the diagnosis of patients and to eventually improve patient care. |
Inácio, Daniel; Amado, Tiago; Pamplona, Ana; Sobral, Daniel; Cunha, Carolina; Santos, Rita F.; Oliveira, Liliana; Rouquié, Nelly; Carmo, Alexandre M.; Lesourne, Renaud; Gomes, Anita Q.; Silva-Santos, Bruno Signature cytokine-associated transcriptome analysis of effector γδ T cells identifies subset-specific regulators of peripheral activation Journal Article In: Nature Immunology, vol. 26, no. 3, pp. 497–510, 2025, ISSN: 1529-2916. @article{inacio_signature_2025,
title = {Signature cytokine-associated transcriptome analysis of effector γδ T cells identifies subset-specific regulators of peripheral activation},
author = {Daniel Inácio and Tiago Amado and Ana Pamplona and Daniel Sobral and Carolina Cunha and Rita F. Santos and Liliana Oliveira and Nelly Rouquié and Alexandre M. Carmo and Renaud Lesourne and Anita Q. Gomes and Bruno Silva-Santos},
doi = {10.1038/s41590-024-02073-8},
issn = {1529-2916},
year = {2025},
date = {2025-03-01},
urldate = {2025-03-01},
journal = {Nature Immunology},
volume = {26},
number = {3},
pages = {497–510},
abstract = {γδ T cells producing either interleukin-17A (γδ17 cells) or interferon-γ (γδIFN cells) are generated in the mouse thymus, but the molecular regulators of their peripheral functions are not fully characterized. Here we established an Il17a-GFP:Ifng-YFP double-reporter mouse strain to analyze at unprecedented depth the transcriptomes of pure γδ17 cell versus γδIFN cell populations from peripheral lymph nodes. Within a very high fraction of differentially expressed genes, we identify a panel of 20 new signature genes in steady-state γδ17 cells versus γδIFN cells, which we further validate in models of experimental autoimmune encephalomyelitis and cerebral malaria, respectively. Among the signature genes, we show that the co-receptor CD6 and the signaling protein Themis promote the activation and proliferation of peripheral γδIFN cells in response to T cell antigen receptor stimulation in vitro and to Plasmodium infection in vivo. This resource can help to understand the distinct activities of effector γδ T cell subsets in pathophysiology.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
γδ T cells producing either interleukin-17A (γδ17 cells) or interferon-γ (γδIFN cells) are generated in the mouse thymus, but the molecular regulators of their peripheral functions are not fully characterized. Here we established an Il17a-GFP:Ifng-YFP double-reporter mouse strain to analyze at unprecedented depth the transcriptomes of pure γδ17 cell versus γδIFN cell populations from peripheral lymph nodes. Within a very high fraction of differentially expressed genes, we identify a panel of 20 new signature genes in steady-state γδ17 cells versus γδIFN cells, which we further validate in models of experimental autoimmune encephalomyelitis and cerebral malaria, respectively. Among the signature genes, we show that the co-receptor CD6 and the signaling protein Themis promote the activation and proliferation of peripheral γδIFN cells in response to T cell antigen receptor stimulation in vitro and to Plasmodium infection in vivo. This resource can help to understand the distinct activities of effector γδ T cell subsets in pathophysiology. |
Lutz, Pierre G.; Lamsoul, Isabelle [Filamins, T helper 2 lymphocytes and asthma : A matter of balance] Journal Article In: Medecine Sciences: M/S, vol. 41, no. 3, pp. 232–235, 2025, ISSN: 1958-5381. @article{lutz_filamins_2025b,
title = {[Filamins, T helper 2 lymphocytes and asthma : A matter of balance]},
author = {Pierre G. Lutz and Isabelle Lamsoul},
doi = {10.1051/medsci/2025032},
issn = {1958-5381},
year = {2025},
date = {2025-03-01},
urldate = {2025-03-01},
journal = {Medecine Sciences: M/S},
volume = {41},
number = {3},
pages = {232–235},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
|
Mühlgrabner, Vanessa; Plach, Angelika; Holler, Johannes; Leitner, Judith; Steinberger, Peter; Dupré, Loïc; Göhring, Janett; Huppa, Johannes B. Gauging antigen recognition by human primary T-cells featuring orthotopically exchanged TCRs of choice Journal Article In: Methods in Cell Biology, vol. 193, pp. 127–154, 2025, ISSN: 0091-679X. @article{muhlgrabner_gauging_2025,
title = {Gauging antigen recognition by human primary T-cells featuring orthotopically exchanged TCRs of choice},
author = {Vanessa Mühlgrabner and Angelika Plach and Johannes Holler and Judith Leitner and Peter Steinberger and Loïc Dupré and Janett Göhring and Johannes B. Huppa},
doi = {10.1016/bs.mcb.2024.03.003},
issn = {0091-679X},
year = {2025},
date = {2025-01-01},
urldate = {2025-01-01},
journal = {Methods in Cell Biology},
volume = {193},
pages = {127–154},
abstract = {Understanding human T-cell antigen recognition in health and disease is becoming increasingly instrumental for monitoring T-cell responses to pathogen challenge and for the rational design of T-cell-based therapies targeting cancer, autoimmunity and organ transplant rejection. Here we showcase a quantitative imaging platform which is based on the use of planar glass-supported lipid bilayers (SLBs). The latter are functionalized with antigen (peptide-loaded HLA) as adhesion and costimulatory molecules (ICAM-1, B7-1) to serve as surrogate antigen presenting cell for antigen recognition by T-cells, which are equipped with T-cell antigen receptors (TCRs) sequenced from antigen-specific patient T-cells. We outline in detail, how the experimental use of SLBs supports recoding and analysis of synaptic antigen engagement and calcium signaling at the single cell level in response to user-defined antigen densities for quantitative comparison.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
Understanding human T-cell antigen recognition in health and disease is becoming increasingly instrumental for monitoring T-cell responses to pathogen challenge and for the rational design of T-cell-based therapies targeting cancer, autoimmunity and organ transplant rejection. Here we showcase a quantitative imaging platform which is based on the use of planar glass-supported lipid bilayers (SLBs). The latter are functionalized with antigen (peptide-loaded HLA) as adhesion and costimulatory molecules (ICAM-1, B7-1) to serve as surrogate antigen presenting cell for antigen recognition by T-cells, which are equipped with T-cell antigen receptors (TCRs) sequenced from antigen-specific patient T-cells. We outline in detail, how the experimental use of SLBs supports recoding and analysis of synaptic antigen engagement and calcium signaling at the single cell level in response to user-defined antigen densities for quantitative comparison. |
2024
|
Maire, Kilian; Chamy, Léa; Ghazali, Samira; Carratala-Lasserre, Manon; Zahm, Margot; Bouisset, Clément; Métais, Arnaud; Combes-Soia, Lucie; Fuente-Vizuete, Lidia; Trad, Hussein; Chaubet, Adeline; Savignac, Magali; de Peredo, Anne Gonzalez; Subramaniam, Arun; Joffre, Olivier; Lutz, Pierre G.; Lamsoul, Isabelle Fine-tuning levels of filamins a and b as a specific mechanism sustaining Th2 lymphocyte functions Journal Article In: Nature Communications, vol. 15, no. 1, pp. 10574, 2024, ISSN: 2041-1723. @article{maire_fine-tuning_2024,
title = {Fine-tuning levels of filamins a and b as a specific mechanism sustaining Th2 lymphocyte functions},
author = {Kilian Maire and Léa Chamy and Samira Ghazali and Manon Carratala-Lasserre and Margot Zahm and Clément Bouisset and Arnaud Métais and Lucie Combes-Soia and Lidia Fuente-Vizuete and Hussein Trad and Adeline Chaubet and Magali Savignac and Anne Gonzalez de Peredo and Arun Subramaniam and Olivier Joffre and Pierre G. Lutz and Isabelle Lamsoul},
doi = {10.1038/s41467-024-53768-3},
issn = {2041-1723},
year = {2024},
date = {2024-12-01},
urldate = {2024-12-01},
journal = {Nature Communications},
volume = {15},
number = {1},
pages = {10574},
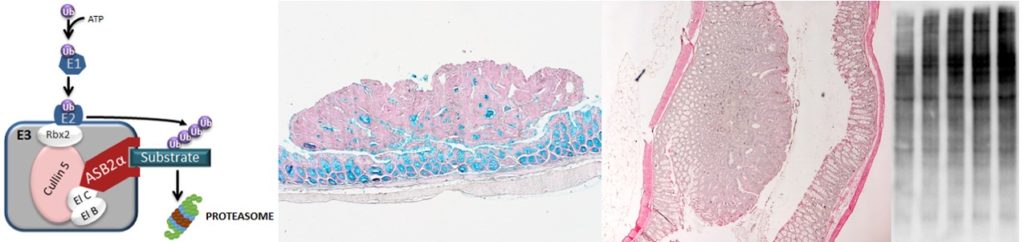
abstract = {Augmenting the portfolio of therapeutics for type 2-driven diseases is crucial to address unmet clinical needs and to design personalized treatment schemes. An attractive therapy for such diseases would consist in targeting the recruitment of T helper 2 (Th2) lymphocytes to inflammatory sites. Herein, we show the degradation of filamins (FLN) a and b by the ASB2α E3 ubiquitin ligase as a mechanism sustaining Th2 lymphocyte functions. Low levels of FLNa and FLNb confer an elongated shape to Th2 lymphocytes associated with efficient αVβ3 integrin-dependent cell migration. Genes encoding the αVβ3 integrin and ASB2α belong to the core of Th2-specific genes. Using genetically modified mice, we find that increasing the levels of FLNa and FLNb in Th2 lymphocytes reduces airway inflammation through diminished Th2 lymphocyte recruitment in inflamed lungs. Collectively, our results highlight ASB2α and its substrates FLNa and FLNb to alter Th2 lymphocyte-mediated responses.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
Augmenting the portfolio of therapeutics for type 2-driven diseases is crucial to address unmet clinical needs and to design personalized treatment schemes. An attractive therapy for such diseases would consist in targeting the recruitment of T helper 2 (Th2) lymphocytes to inflammatory sites. Herein, we show the degradation of filamins (FLN) a and b by the ASB2α E3 ubiquitin ligase as a mechanism sustaining Th2 lymphocyte functions. Low levels of FLNa and FLNb confer an elongated shape to Th2 lymphocytes associated with efficient αVβ3 integrin-dependent cell migration. Genes encoding the αVβ3 integrin and ASB2α belong to the core of Th2-specific genes. Using genetically modified mice, we find that increasing the levels of FLNa and FLNb in Th2 lymphocytes reduces airway inflammation through diminished Th2 lymphocyte recruitment in inflamed lungs. Collectively, our results highlight ASB2α and its substrates FLNa and FLNb to alter Th2 lymphocyte-mediated responses. |
Cadot, Sarah; Audebert, Chloe; Dion, Charlotte; Ken, Soleakhena; Dupré, Loic; Largeaud, Laetitia; Laurent, Camille; Ysebaert, Loic; Crauste, Fabien; Quillet-Mary, Anne New pharmacodynamic parameters linked with ibrutinib responses in chronic lymphocytic leukemia: Prospective study in real-world patients and mathematical modeling Journal Article In: PLoS medicine, vol. 21, no. 7, pp. e1004430, 2024, ISSN: 1549-1676. @article{cadot_new_2024,
title = {New pharmacodynamic parameters linked with ibrutinib responses in chronic lymphocytic leukemia: Prospective study in real-world patients and mathematical modeling},
author = {Sarah Cadot and Chloe Audebert and Charlotte Dion and Soleakhena Ken and Loic Dupré and Laetitia Largeaud and Camille Laurent and Loic Ysebaert and Fabien Crauste and Anne Quillet-Mary},
doi = {10.1371/journal.pmed.1004430},
issn = {1549-1676},
year = {2024},
date = {2024-07-01},
urldate = {2024-07-01},
journal = {PLoS medicine},
volume = {21},
number = {7},
pages = {e1004430},
abstract = {BACKGROUND: One of the first clinical observations of ibrutinib activity in the treatment of chronic lymphocytic leukemia (CLL) is a rapid decline in lymph nodes size. This phenomenon is accompanied by an hyperlymphocytosis, either transient or prolonged, which is associated with distinct clinical responses and thus has an impact on long-term outcomes. Understanding which factors determine distinct disease courses upon ibrutinib treatment remains a scientific challenge. METHODS AND FINDINGS: From 2016 to 2021, we conducted a longitudinal and observational study in 2 cohorts of patients with chronic lymphocytic leukemia (CLL) (cohort 1},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
BACKGROUND: One of the first clinical observations of ibrutinib activity in the treatment of chronic lymphocytic leukemia (CLL) is a rapid decline in lymph nodes size. This phenomenon is accompanied by an hyperlymphocytosis, either transient or prolonged, which is associated with distinct clinical responses and thus has an impact on long-term outcomes. Understanding which factors determine distinct disease courses upon ibrutinib treatment remains a scientific challenge. METHODS AND FINDINGS: From 2016 to 2021, we conducted a longitudinal and observational study in 2 cohorts of patients with chronic lymphocytic leukemia (CLL) (cohort 1 |
Fusaro, Mathieu; Dupré, Loïc Mechanisms underlying skin inflammation of DOCK8 deficiency Journal Article In: The Journal of Allergy and Clinical Immunology, vol. 154, no. 1, pp. 88–90, 2024, ISSN: 1097-6825. @article{fusaro_mechanisms_2024,
title = {Mechanisms underlying skin inflammation of DOCK8 deficiency},
author = {Mathieu Fusaro and Loïc Dupré},
doi = {10.1016/j.jaci.2024.04.026},
issn = {1097-6825},
year = {2024},
date = {2024-07-01},
urldate = {2024-07-01},
journal = {The Journal of Allergy and Clinical Immunology},
volume = {154},
number = {1},
pages = {88–90},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
|
Mélique, Suzanne; Vadel, Aurélie; Rouquié, Nelly; Yang, Cui; Bories, Cyrielle; Cotineau, Coline; Saoudi, Abdelhadi; Fazilleau, Nicolas; Lesourne, Renaud THEMIS promotes T cell development and maintenance by rising the signaling threshold of the inhibitory receptor BTLA Journal Article In: Proc Natl Acad Sci U S A, vol. 121, no. 20, pp. e2318773121, 2024, ISSN: 1091-6490. @article{pmid38713628,
title = {THEMIS promotes T cell development and maintenance by rising the signaling threshold of the inhibitory receptor BTLA},
author = {Suzanne Mélique and Aurélie Vadel and Nelly Rouquié and Cui Yang and Cyrielle Bories and Coline Cotineau and Abdelhadi Saoudi and Nicolas Fazilleau and Renaud Lesourne},
doi = {10.1073/pnas.2318773121},
issn = {1091-6490},
year = {2024},
date = {2024-05-01},
urldate = {2024-05-01},
journal = {Proc Natl Acad Sci U S A},
volume = {121},
number = {20},
pages = {e2318773121},
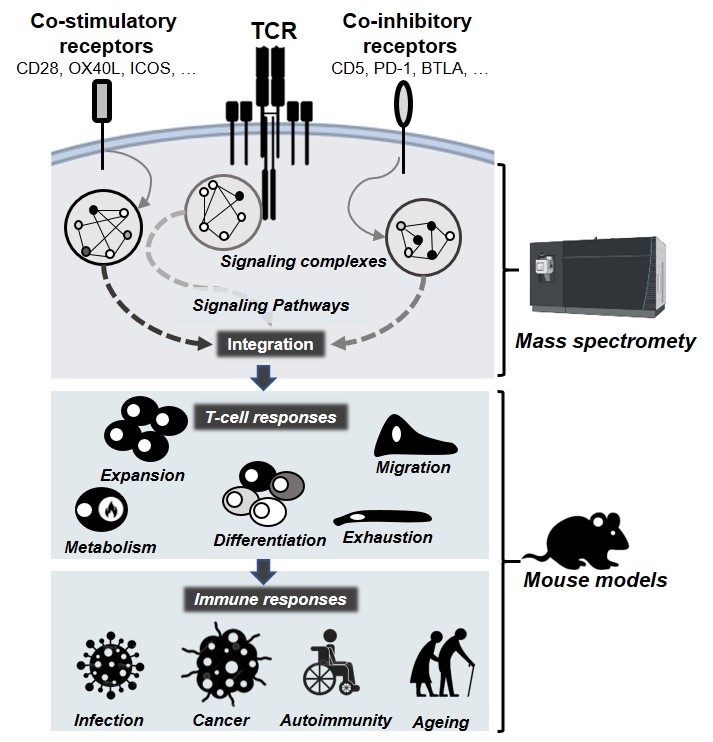
abstract = {The current paradigm about the function of T cell immune checkpoints is that these receptors switch on inhibitory signals upon cognate ligand interaction. We here revisit this simple switch model and provide evidence that the T cell lineage protein THEMIS enhances the signaling threshold at which the immune checkpoint BTLA (B- and T-lymphocyte attenuator) represses T cell responses. THEMIS is recruited to the cytoplasmic domain of BTLA and blocks its signaling capacity by promoting/stabilizing the oxidation of the catalytic cysteine of the tyrosine phosphatase SHP-1. In contrast, THEMIS has no detectable effect on signaling pathways regulated by PD-1 (Programmed cell death protein 1), which depend mainly on the tyrosine phosphatase SHP-2. BTLA inhibitory signaling is tuned according to the THEMIS expression level, making CD8+ T cells more resistant to BTLA-mediated inhibition than CD4+ T cells. In the absence of THEMIS, the signaling capacity of BTLA is exacerbated, which results in the attenuation of signals driven by the T cell antigen receptor and by receptors for IL-2 and IL-15, consequently hampering thymocyte positive selection and peripheral CD8+ T cell maintenance. By characterizing the pivotal role of THEMIS in restricting the transmission of BTLA signals, our study suggests that immune checkpoint operability is conditioned by intracellular signal attenuators.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
The current paradigm about the function of T cell immune checkpoints is that these receptors switch on inhibitory signals upon cognate ligand interaction. We here revisit this simple switch model and provide evidence that the T cell lineage protein THEMIS enhances the signaling threshold at which the immune checkpoint BTLA (B- and T-lymphocyte attenuator) represses T cell responses. THEMIS is recruited to the cytoplasmic domain of BTLA and blocks its signaling capacity by promoting/stabilizing the oxidation of the catalytic cysteine of the tyrosine phosphatase SHP-1. In contrast, THEMIS has no detectable effect on signaling pathways regulated by PD-1 (Programmed cell death protein 1), which depend mainly on the tyrosine phosphatase SHP-2. BTLA inhibitory signaling is tuned according to the THEMIS expression level, making CD8+ T cells more resistant to BTLA-mediated inhibition than CD4+ T cells. In the absence of THEMIS, the signaling capacity of BTLA is exacerbated, which results in the attenuation of signals driven by the T cell antigen receptor and by receptors for IL-2 and IL-15, consequently hampering thymocyte positive selection and peripheral CD8+ T cell maintenance. By characterizing the pivotal role of THEMIS in restricting the transmission of BTLA signals, our study suggests that immune checkpoint operability is conditioned by intracellular signal attenuators. |
Marrocco, Remi; Bernard, Isabelle; Joulia, Emeline; Barascud, Rebecca; Dejean, Anne S.; Lesourne, Renaud; Saoudi, Abdelhadi Positive regulation of Vav1 by Themis controls CD4 T cell pathogenicity in a mouse model of central nervous system inflammation Journal Article In: Cellular and molecular life sciences: CMLS, vol. 81, no. 1, pp. 161, 2024, ISSN: 1420-9071. @article{marrocco_positive_2024,
title = {Positive regulation of Vav1 by Themis controls CD4 T cell pathogenicity in a mouse model of central nervous system inflammation},
author = {Remi Marrocco and Isabelle Bernard and Emeline Joulia and Rebecca Barascud and Anne S. Dejean and Renaud Lesourne and Abdelhadi Saoudi},
doi = {10.1007/s00018-024-05203-5},
issn = {1420-9071},
year = {2024},
date = {2024-04-01},
urldate = {2024-04-01},
journal = {Cellular and molecular life sciences: CMLS},
volume = {81},
number = {1},
pages = {161},
abstract = {The susceptibility to autoimmune diseases is conditioned by the association of modest genetic alterations which altogether weaken self-tolerance. The mechanism whereby these genetic interactions modulate T-cell pathogenicity remains largely uncovered. Here, we investigated the epistatic interaction of two interacting proteins involved in T Cell Receptor signaling and which were previously associated with the development of Multiple Sclerosis. To this aim, we used mice expressing an hypomorphic variant of Vav1 (Vav1R63W), combined with a T cell-conditional deletion of Themis. We show that the combined mutations in Vav1 and Themis induce a strong attenuation of the severity of Experimental Autoimmune Encephalomyelitis (EAE), contrasting with the moderate effect of the single mutation in each of those two proteins. This genotype-dependent gradual decrease of EAE severity correlates with decreased quantity of phosphorylated Vav1 in CD4 T cells, establishing that Themis promotes the development of encephalitogenic Tconv response by enhancing Vav1 activity. We also show that the cooperative effect of Themis and Vav1 on EAE severity is independent of regulatory T cells and unrelated to the impact of Themis on thymic selection. Rather, it results from decreased production of pro-inflammatory cytokines (IFN-γ, IL-17, TNF and GM-CSF) and reduced T cell infiltration in the CNS. Together, our results provide a rationale to study combination of related genes, in addition to single gene association, to better understand the genetic bases of human diseases.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
The susceptibility to autoimmune diseases is conditioned by the association of modest genetic alterations which altogether weaken self-tolerance. The mechanism whereby these genetic interactions modulate T-cell pathogenicity remains largely uncovered. Here, we investigated the epistatic interaction of two interacting proteins involved in T Cell Receptor signaling and which were previously associated with the development of Multiple Sclerosis. To this aim, we used mice expressing an hypomorphic variant of Vav1 (Vav1R63W), combined with a T cell-conditional deletion of Themis. We show that the combined mutations in Vav1 and Themis induce a strong attenuation of the severity of Experimental Autoimmune Encephalomyelitis (EAE), contrasting with the moderate effect of the single mutation in each of those two proteins. This genotype-dependent gradual decrease of EAE severity correlates with decreased quantity of phosphorylated Vav1 in CD4 T cells, establishing that Themis promotes the development of encephalitogenic Tconv response by enhancing Vav1 activity. We also show that the cooperative effect of Themis and Vav1 on EAE severity is independent of regulatory T cells and unrelated to the impact of Themis on thymic selection. Rather, it results from decreased production of pro-inflammatory cytokines (IFN-γ, IL-17, TNF and GM-CSF) and reduced T cell infiltration in the CNS. Together, our results provide a rationale to study combination of related genes, in addition to single gene association, to better understand the genetic bases of human diseases. |
Rey-Barroso, Javier; Munaretto, Alice; Rouquié, Nelly; Mougel, Aurélie; Chassan, Malika; Gadat, Sébastien; Dewingle, Océane; Poincloux, Renaud; Cadot, Sarah; Ysebaert, Loïc; Quillet-Mary, Anne; Dupré, Loïc Lymphocyte migration and retention properties affected by ibrutinib in chronic lymphocytic leukemia Journal Article In: Haematologica, vol. 109, no. 3, pp. 809–823, 2024, ISSN: 1592-8721. @article{rey-barroso_lymphocyte_2024,
title = {Lymphocyte migration and retention properties affected by ibrutinib in chronic lymphocytic leukemia},
author = {Javier Rey-Barroso and Alice Munaretto and Nelly Rouquié and Aurélie Mougel and Malika Chassan and Sébastien Gadat and Océane Dewingle and Renaud Poincloux and Sarah Cadot and Loïc Ysebaert and Anne Quillet-Mary and Loïc Dupré},
doi = {10.3324/haematol.2022.282466},
issn = {1592-8721},
year = {2024},
date = {2024-03-01},
urldate = {2024-03-01},
journal = {Haematologica},
volume = {109},
number = {3},
pages = {809–823},
abstract = {The Bruton tyrosine kinase (BTK) inhibitor ibrutinib is widely used for treatment of patients with relapsed/refractory or treatment-naïve chronic lymphocytic leukemia (CLL). A prominent effect of ibrutinib is to disrupt the retention of CLL cells from supportive lymphoid tissues, by altering BTK-dependent adhesion and migration. To further explore the mechanism of action of ibrutinib and its potential impact on non-leukemic cells, we quantified multiple motility and adhesion parameters of human primary CLL cells and non-leukemic lymphoid cells. In vitro, ibrutinib affected CCL19-, CXCL12- and CXCL13-evoked migration behavior of CLL cells and non-neoplastic lymphocytes, by reducing both motility speed and directionality. De-phosphorylation of BTK induced by ibrutinib in CLL cells was associated with defective polarization over fibronectin and inability to assemble the immunological synapse upon B-cell receptor engagement. In patients' samples collected during a 6-month monitoring of therapy, chemokine-evoked migration was repressed in CLL cells and marginally reduced in T cells. This was accompanied by profound modulation of the expression of chemokine receptors and adhesion molecules. Remarkably, the relative expression of the receptors governing lymph node entry (CCR7) versus exit (S1PR1) stood out as a reliable predictive marker of the clinically relevant treatment-induced lymphocytosis. Together, our data reveal a multifaceted modulation of motility and adhesive properties of ibrutinib on both CLL leukemic cell and T-cell populations and point to intrinsic differences in CLL recirculation properties as an underlying cause for variability in treatment response.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
The Bruton tyrosine kinase (BTK) inhibitor ibrutinib is widely used for treatment of patients with relapsed/refractory or treatment-naïve chronic lymphocytic leukemia (CLL). A prominent effect of ibrutinib is to disrupt the retention of CLL cells from supportive lymphoid tissues, by altering BTK-dependent adhesion and migration. To further explore the mechanism of action of ibrutinib and its potential impact on non-leukemic cells, we quantified multiple motility and adhesion parameters of human primary CLL cells and non-leukemic lymphoid cells. In vitro, ibrutinib affected CCL19-, CXCL12- and CXCL13-evoked migration behavior of CLL cells and non-neoplastic lymphocytes, by reducing both motility speed and directionality. De-phosphorylation of BTK induced by ibrutinib in CLL cells was associated with defective polarization over fibronectin and inability to assemble the immunological synapse upon B-cell receptor engagement. In patients' samples collected during a 6-month monitoring of therapy, chemokine-evoked migration was repressed in CLL cells and marginally reduced in T cells. This was accompanied by profound modulation of the expression of chemokine receptors and adhesion molecules. Remarkably, the relative expression of the receptors governing lymph node entry (CCR7) versus exit (S1PR1) stood out as a reliable predictive marker of the clinically relevant treatment-induced lymphocytosis. Together, our data reveal a multifaceted modulation of motility and adhesive properties of ibrutinib on both CLL leukemic cell and T-cell populations and point to intrinsic differences in CLL recirculation properties as an underlying cause for variability in treatment response. |
Kamnev, Anton; Mehta, Tanvi; Wielscher, Matthias; Chaves, Beatriz; Lacouture, Claire; Mautner, Anna-Katharina; Shaw, Lisa E.; Caldera, Michael; Menche, Jörg; Weninger, Wolfgang P.; Farlik, Matthias; Boztug, Kaan; Dupré, Loïc Coordinated ARP2/3 and glycolytic activities regulate the morphological and functional fitness of human CD8+ T cells Journal Article In: Cell Reports, vol. 43, no. 3, pp. 113853, 2024, ISSN: 2211-1247. @article{kamnev_coordinated_2024,
title = {Coordinated ARP2/3 and glycolytic activities regulate the morphological and functional fitness of human CD8+ T cells},
author = {Anton Kamnev and Tanvi Mehta and Matthias Wielscher and Beatriz Chaves and Claire Lacouture and Anna-Katharina Mautner and Lisa E. Shaw and Michael Caldera and Jörg Menche and Wolfgang P. Weninger and Matthias Farlik and Kaan Boztug and Loïc Dupré},
doi = {10.1016/j.celrep.2024.113853},
issn = {2211-1247},
year = {2024},
date = {2024-03-01},
urldate = {2024-03-01},
journal = {Cell Reports},
volume = {43},
number = {3},
pages = {113853},
abstract = {Actin cytoskeleton remodeling sustains the ability of cytotoxic T cells to search for target cells and eliminate them. We here investigated the relationship between energetic status, actin remodeling, and functional fitness in human CD8+ effector T cells. Cell spreading during migration or immunological synapse assembly mirrored cytotoxic activity. Morphological and functional fitness were boosted by interleukin-2 (IL-2), which also stimulated the transcription of glycolytic enzymes, actin isoforms, and actin-related protein (ARP)2/3 complex subunits. This molecular program scaled with F-actin content and cell spreading. Inhibiting glycolysis impaired F-actin remodeling at the lamellipodium, chemokine-driven motility, and adhesion, while mitochondrial oxidative phosphorylation blockade impacted cell elongation during confined migration. The severe morphological and functional defects of ARPC1B-deficient T cells were only partially corrected by IL-2, emphasizing ARP2/3-mediated actin polymerization as a crucial energy state integrator. The study therefore underscores the tight coordination between metabolic and actin remodeling programs to sustain the cytotoxic activity of CD8+ T cells.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
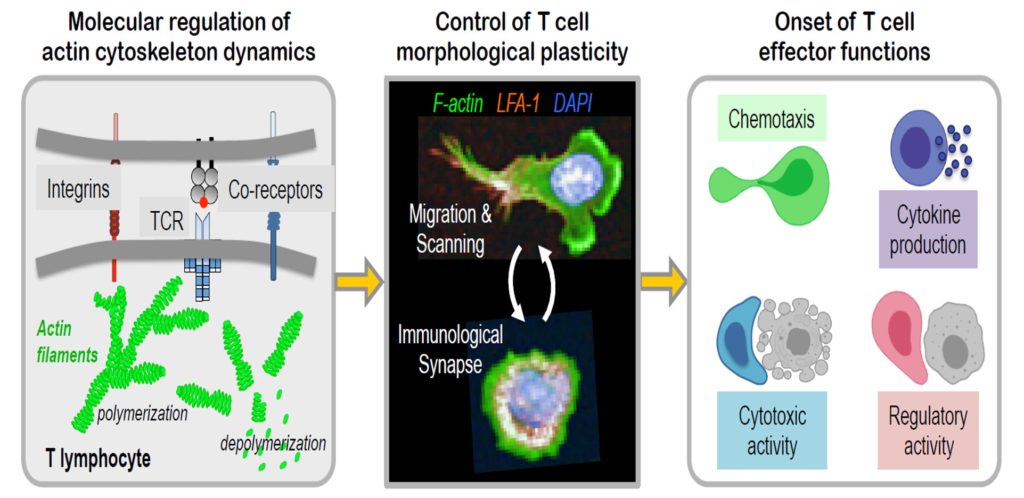
Actin cytoskeleton remodeling sustains the ability of cytotoxic T cells to search for target cells and eliminate them. We here investigated the relationship between energetic status, actin remodeling, and functional fitness in human CD8+ effector T cells. Cell spreading during migration or immunological synapse assembly mirrored cytotoxic activity. Morphological and functional fitness were boosted by interleukin-2 (IL-2), which also stimulated the transcription of glycolytic enzymes, actin isoforms, and actin-related protein (ARP)2/3 complex subunits. This molecular program scaled with F-actin content and cell spreading. Inhibiting glycolysis impaired F-actin remodeling at the lamellipodium, chemokine-driven motility, and adhesion, while mitochondrial oxidative phosphorylation blockade impacted cell elongation during confined migration. The severe morphological and functional defects of ARPC1B-deficient T cells were only partially corrected by IL-2, emphasizing ARP2/3-mediated actin polymerization as a crucial energy state integrator. The study therefore underscores the tight coordination between metabolic and actin remodeling programs to sustain the cytotoxic activity of CD8+ T cells. |
Lacouture, Claire; Chaves, Beatriz; Guipouy, Delphine; Houmadi, Raïssa; Duplan-Eche, Valérie; Allart, Sophie; Destainville, Nicolas; Dupré, Loïc LFA-1 nanoclusters integrate TCR stimulation strength to tune T-cell cytotoxic activity Journal Article In: Nature Communications, vol. 15, no. 1, pp. 407, 2024, ISSN: 2041-1723, (Number: 1
Publisher: Nature Publishing Group). @article{lacouture_lfa-1_2024,
title = {LFA-1 nanoclusters integrate TCR stimulation strength to tune T-cell cytotoxic activity},
author = {Claire Lacouture and Beatriz Chaves and Delphine Guipouy and Raïssa Houmadi and Valérie Duplan-Eche and Sophie Allart and Nicolas Destainville and Loïc Dupré},
url = {https://www.nature.com/articles/s41467-024-44688-3},
doi = {10.1038/s41467-024-44688-3},
issn = {2041-1723},
year = {2024},
date = {2024-01-01},
urldate = {2024-01-01},
journal = {Nature Communications},
volume = {15},
number = {1},
pages = {407},
abstract = {T-cell cytotoxic function relies on the cooperation between the highly specific but poorly adhesive T-cell receptor (TCR) and the integrin LFA-1. How LFA-1-mediated adhesion may scale with TCR stimulation strength is ill-defined. Here, we show that LFA-1 conformation activation scales with TCR stimulation to calibrate human T-cell cytotoxicity. Super-resolution microscopy analysis reveals that >1000 LFA-1 nanoclusters provide a discretized platform at the immunological synapse to translate TCR engagement and density of the LFA-1 ligand ICAM-1 into graded adhesion. Indeed, the number of high-affinity conformation LFA-1 nanoclusters increases as a function of TCR triggering strength. Blockade of LFA-1 conformational activation impairs adhesion to target cells and killing. However, it occurs at a lower TCR stimulation threshold than lytic granule exocytosis implying that it licenses, rather than directly controls, the killing decision. We conclude that the organization of LFA-1 into nanoclusters provides a calibrated system to adjust T-cell killing to the antigen stimulation strength.},
note = {Number: 1
Publisher: Nature Publishing Group},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
T-cell cytotoxic function relies on the cooperation between the highly specific but poorly adhesive T-cell receptor (TCR) and the integrin LFA-1. How LFA-1-mediated adhesion may scale with TCR stimulation strength is ill-defined. Here, we show that LFA-1 conformation activation scales with TCR stimulation to calibrate human T-cell cytotoxicity. Super-resolution microscopy analysis reveals that >1000 LFA-1 nanoclusters provide a discretized platform at the immunological synapse to translate TCR engagement and density of the LFA-1 ligand ICAM-1 into graded adhesion. Indeed, the number of high-affinity conformation LFA-1 nanoclusters increases as a function of TCR triggering strength. Blockade of LFA-1 conformational activation impairs adhesion to target cells and killing. However, it occurs at a lower TCR stimulation threshold than lytic granule exocytosis implying that it licenses, rather than directly controls, the killing decision. We conclude that the organization of LFA-1 into nanoclusters provides a calibrated system to adjust T-cell killing to the antigen stimulation strength. |
2023
|
Block, Jana; Rashkova, Christina; Castanon, Irinka; Zoghi, Samaneh; Platon, Jessica; Ardy, Rico C.; Fujiwara, Mitsuhiro; Chaves, Beatriz; Schoppmeyer, Rouven; van der Made, Caspar I.; Jimenez Heredia, Raul; Harms, Frederike L.; Alavi, Samin; Alsina, Laia; Sanchez Moreno, Paula; Ávila Polo, Rainiero; Cabrera-Pérez, Rocío; Kostel Bal, Sevgi; Pfajfer, Laurène; Ransmayr, Bernhard; Mautner, Anna-Katharina; Kondo, Ryohei; Tinnacher, Anna; Caldera, Michael; Schuster, Michael; Domínguez Conde, Cecilia; Platzer, René; Salzer, Elisabeth; Boyer, Thomas; Brunner, Han G.; Nooitgedagt-Frons, Judith E.; Iglesias, Estíbaliz; Deyà-Martinez, Angela; Camacho-Lovillo, Marisol; Menche, Jörg; Bock, Christoph; Huppa, Johannes B.; Pickl, Winfried F.; Distel, Martin; Yoder, Jeffrey A.; Traver, David; Engelhardt, Karin R.; Linden, Tobias; Kager, Leo; Hannich, J. Thomas; Hoischen, Alexander; Hambleton, Sophie; Illsinger, Sabine; Da Costa, Lydie; Kutsche, Kerstin; Chavoshzadeh, Zahra; van Buul, Jaap D.; Antón, Jordi; Calzada-Hernández, Joan; Neth, Olaf; Viaud, Julien; Nishikimi, Akihiko; Dupré, Loïc; Boztug, Kaan Systemic Inflammation and Normocytic Anemia in DOCK11 Deficiency Journal Article In: N Engl J Med, 2023, ISSN: 1533-4406. @article{block_systemic_2023,
title = {Systemic Inflammation and Normocytic Anemia in DOCK11 Deficiency},
author = {Block, Jana and Rashkova, Christina and Castanon, Irinka and Zoghi, Samaneh and Platon, Jessica and Ardy, Rico C. and Fujiwara, Mitsuhiro and Chaves, Beatriz and Schoppmeyer, Rouven and van der Made, Caspar I. and Jimenez Heredia, Raul and Harms, Frederike L. and Alavi, Samin and Alsina, Laia and Sanchez Moreno, Paula and Ávila Polo, Rainiero and Cabrera-Pérez, Rocío and Kostel Bal, Sevgi and Pfajfer, Laurène and Ransmayr, Bernhard and Mautner, Anna-Katharina and Kondo, Ryohei and Tinnacher, Anna and Caldera, Michael and Schuster, Michael and Domínguez Conde, Cecilia and Platzer, René and Salzer, Elisabeth and Boyer, Thomas and Brunner, Han G. and Nooitgedagt-Frons, Judith E. and Iglesias, Estíbaliz and Deyà-Martinez, Angela and Camacho-Lovillo, Marisol and Menche, Jörg and Bock, Christoph and Huppa, Johannes B. and Pickl, Winfried F. and Distel, Martin and Yoder, Jeffrey A. and Traver, David and Engelhardt, Karin R. and Linden, Tobias and Kager, Leo and Hannich, J. Thomas and Hoischen, Alexander and Hambleton, Sophie and Illsinger, Sabine and Da Costa, Lydie and Kutsche, Kerstin and Chavoshzadeh, Zahra and van Buul, Jaap D. and Antón, Jordi and Calzada-Hernández, Joan and Neth, Olaf and Viaud, Julien and Nishikimi, Akihiko and Dupré, Loïc and Boztug, Kaan},
doi = {10.1056/NEJMoa2210054},
issn = {1533-4406},
year = {2023},
date = {2023-06-01},
journal = {N Engl J Med},
abstract = {BACKGROUND: Increasing evidence links genetic defects affecting actin-regulatory proteins to diseases with severe autoimmunity and autoinflammation, yet the underlying molecular mechanisms are poorly understood. Dedicator of cytokinesis 11 (DOCK11) activates the small Rho guanosine triphosphatase (GTPase) cell division cycle 42 (CDC42), a central regulator of actin cytoskeleton dynamics. The role of DOCK11 in human immune-cell function and disease remains unknown.
METHODS: We conducted genetic, immunologic, and molecular assays in four patients from four unrelated families who presented with infections, early-onset severe immune dysregulation, normocytic anemia of variable severity associated with anisopoikilocytosis, and developmental delay. Functional assays were performed in patient-derived cells, as well as in mouse and zebrafish models.
RESULTS: We identified rare, X-linked germline mutations in DOCK11 in the patients, leading to a loss of protein expression in two patients and impaired CDC42 activation in all four patients. Patient-derived T cells did not form filopodia and showed abnormal migration. In addition, the patient-derived T cells, as well as the T cells from Dock11-knockout mice, showed overt activation and production of proinflammatory cytokines that were associated with an increased degree of nuclear translocation of nuclear factor of activated T cell 1 (NFATc1). Anemia and aberrant erythrocyte morphologic features were recapitulated in a newly generated dock11-knockout zebrafish model, and anemia was amenable to rescue on ectopic expression of constitutively active CDC42.
CONCLUSIONS: Germline hemizygous loss-of-function mutations affecting the actin regulator DOCK11 were shown to cause a previously unknown inborn error of hematopoiesis and immunity characterized by severe immune dysregulation and systemic inflammation, recurrent infections, and anemia. (Funded by the European Research Council and others.).},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
BACKGROUND: Increasing evidence links genetic defects affecting actin-regulatory proteins to diseases with severe autoimmunity and autoinflammation, yet the underlying molecular mechanisms are poorly understood. Dedicator of cytokinesis 11 (DOCK11) activates the small Rho guanosine triphosphatase (GTPase) cell division cycle 42 (CDC42), a central regulator of actin cytoskeleton dynamics. The role of DOCK11 in human immune-cell function and disease remains unknown.
METHODS: We conducted genetic, immunologic, and molecular assays in four patients from four unrelated families who presented with infections, early-onset severe immune dysregulation, normocytic anemia of variable severity associated with anisopoikilocytosis, and developmental delay. Functional assays were performed in patient-derived cells, as well as in mouse and zebrafish models.
RESULTS: We identified rare, X-linked germline mutations in DOCK11 in the patients, leading to a loss of protein expression in two patients and impaired CDC42 activation in all four patients. Patient-derived T cells did not form filopodia and showed abnormal migration. In addition, the patient-derived T cells, as well as the T cells from Dock11-knockout mice, showed overt activation and production of proinflammatory cytokines that were associated with an increased degree of nuclear translocation of nuclear factor of activated T cell 1 (NFATc1). Anemia and aberrant erythrocyte morphologic features were recapitulated in a newly generated dock11-knockout zebrafish model, and anemia was amenable to rescue on ectopic expression of constitutively active CDC42.
CONCLUSIONS: Germline hemizygous loss-of-function mutations affecting the actin regulator DOCK11 were shown to cause a previously unknown inborn error of hematopoiesis and immunity characterized by severe immune dysregulation and systemic inflammation, recurrent infections, and anemia. (Funded by the European Research Council and others.). |
Kostel Bal, Sevgi; Giuliani, Sarah; Block, Jana; Repiscak, Peter; Hafemeister, Christoph; Shahin, Tala; Kasap, Nurhan; Ransmayr, Bernhard; Miao, Yirun; van de Wetering, Cheryl; Frohne, Alexandra; Jimenez-Heredia, Raul; Schuster, Michael K.; Zoghi, Samaneh; Hertlein, Vanessa; Thian, Marini; Bykov, Aleksandr; Babayeva, Royala; Bilgic Eltan, Sevgi; Karakoc-Aydiner, Elif; Shaw, Lisa E.; Chowdhury, Iftekhar; Varjosalo, Markku; Argüello, Rafael Jose; Farlik, Matthias; Ozen, Ahmet; Serfling, Edgar Albert Ernst; Dupré, Loïc; Bock, Christoph; Halbritter, Florian; Hannich, J. Thomas; Castanon, Irinka; Kraakman, Michael J.; Baris, Safa; Boztug, Kaan Biallelic NFATC1 mutations cause an inborn error of immunity with impaired CD8+ Ŧ-cell function and perturbed glycolysis Journal Article In: Blood, pp. blood.2022018303, 2023, ISSN: 1528-0020. @article{kostel_bal_biallelic_2023,
title = {Biallelic NFATC1 mutations cause an inborn error of immunity with impaired CD8+ Ŧ-cell function and perturbed glycolysis},
author = {Kostel Bal, Sevgi and Giuliani, Sarah and Block, Jana and Repiscak, Peter and Hafemeister, Christoph and Shahin, Tala and Kasap, Nurhan and Ransmayr, Bernhard and Miao, Yirun and van de Wetering, Cheryl and Frohne, Alexandra and Jimenez-Heredia, Raul and Schuster, Michael K. and Zoghi, Samaneh and Hertlein, Vanessa and Thian, Marini and Bykov, Aleksandr and Babayeva, Royala and Bilgic Eltan, Sevgi and Karakoc-Aydiner, Elif and Shaw, Lisa E. and Chowdhury, Iftekhar and Varjosalo, Markku and Argüello, Rafael Jose and Farlik, Matthias and Ozen, Ahmet and Serfling, Edgar Albert Ernst and Dupré, Loïc and Bock, Christoph and Halbritter, Florian and Hannich, J. Thomas and Castanon, Irinka and Kraakman, Michael J. and Baris, Safa and Boztug, Kaan},
doi = {10.1182/blood.2022018303},
issn = {1528-0020},
year = {2023},
date = {2023-01-01},
journal = {Blood},
pages = {blood.2022018303},
abstract = {The NFAT family of transcription factors plays central roles in adaptive immunity in murine models, however, their contribution to human immune homeostasis remains poorly defined. In a multigenerational pedigree, we identified three patients carrying germline biallelic missense variants in NFATC1, presenting with recurrent infections, hypogammaglobulinemia and decreased antibody responses. The compound heterozygous NFATC1 variants identified in the patients caused decreased stability and reduced binding of DNA and interacting proteins. We observed defects in early activation and proliferation of T and B cells from these patients, amenable to reconstitution upon genetic rescue. Following stimulation, T-cell activation and proliferation were impaired, reaching that of healthy controls with delay indicative of an adaptive capacity of the cells. Assessment of the metabolic capacity of patient T cells, revealed that NFATc1-dysfunction rendered T cells unable to engage in glycolysis following stimulation, although oxidative metabolic processes were intact. We hypothesized that NFATc1-mutant T cells could compensate for the energy deficit due to defective glycolysis by enhanced lipid metabolism as an adaptation, leading to a delayed, but not lost activation responses. Indeed, we observed increased 13C-labelled palmitate incorporation into citrate indicating higher fatty acid oxidation and we demonstrated that metformin and rosiglitazone improved patient T-cell effector functions. Collectively, enabled by our molecular dissection of NFATC1 mutations and extending the role of NFATc1 in human immunity beyond receptor signaling, and reveal evidence of metabolic plasticity in the context of impaired glycolysis observed in patient T cells to remedy delayed effector responses.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
The NFAT family of transcription factors plays central roles in adaptive immunity in murine models, however, their contribution to human immune homeostasis remains poorly defined. In a multigenerational pedigree, we identified three patients carrying germline biallelic missense variants in NFATC1, presenting with recurrent infections, hypogammaglobulinemia and decreased antibody responses. The compound heterozygous NFATC1 variants identified in the patients caused decreased stability and reduced binding of DNA and interacting proteins. We observed defects in early activation and proliferation of T and B cells from these patients, amenable to reconstitution upon genetic rescue. Following stimulation, T-cell activation and proliferation were impaired, reaching that of healthy controls with delay indicative of an adaptive capacity of the cells. Assessment of the metabolic capacity of patient T cells, revealed that NFATc1-dysfunction rendered T cells unable to engage in glycolysis following stimulation, although oxidative metabolic processes were intact. We hypothesized that NFATc1-mutant T cells could compensate for the energy deficit due to defective glycolysis by enhanced lipid metabolism as an adaptation, leading to a delayed, but not lost activation responses. Indeed, we observed increased 13C-labelled palmitate incorporation into citrate indicating higher fatty acid oxidation and we demonstrated that metformin and rosiglitazone improved patient T-cell effector functions. Collectively, enabled by our molecular dissection of NFATC1 mutations and extending the role of NFATc1 in human immunity beyond receptor signaling, and reveal evidence of metabolic plasticity in the context of impaired glycolysis observed in patient T cells to remedy delayed effector responses. |
Dupré, Loïc; Prunier, Guilhèn Deciphering actin remodelling in immune cells through the prism of actin-related inborn errors of immunity Journal Article In: Eur J Cell Biol, vol. 102, no. 1, pp. 151283, 2023, ISSN: 1618-1298. @article{dupre_deciphering_2023,
title = {Deciphering actin remodelling in immune cells through the prism of actin-related inborn errors of immunity},
author = {Dupré, Loïc and Prunier, Guilhèn},
doi = {10.1016/j.ejcb.2022.151283},
issn = {1618-1298},
year = {2023},
date = {2023-01-01},
journal = {Eur J Cell Biol},
volume = {102},
number = {1},
pages = {151283},
abstract = {Actin cytoskeleton remodelling drives cell motility, cell to cell contacts, as well as membrane and organelle dynamics. Those cellular activities operate at a particularly high pace in immune cells since these cells migrate through various tissues, interact with multiple cellular partners, ingest microorganisms and secrete effector molecules. The central and multifaceted role of actin cytoskeleton remodelling in sustaining immune cell tasks in humans is highlighted by rare inborn errors of immunity due to mutations in genes encoding proximal and distal actin regulators. In line with the specificity of some of the actin-based processes at work in immune cells, the expression of some of the affected genes, such as WAS, ARPC1B and HEM1 is restricted to the hematopoietic compartment. Exploration of these natural deficiencies highlights the fact that the molecular control of actin remodelling is tuned distinctly in the various subsets of myeloid and lymphoid immune cells and sustains different networks associated with a vast array of specialized tasks. Furthermore, defects in individual actin remodelling proteins are usually associated with partial cellular impairments highlighting the plasticity of actin cytoskeleton remodelling. This review covers the roles of disease-associated actin regulators in promoting the actin-based processes of immune cells. It focuses on the specific molecular function of those regulators across various immune cell subsets and in response to different stimuli. Given the fact that numerous immune-related actin defects have only been characterized recently, we further discuss the challenges lying ahead to decipher the underlying patho-mechanisms.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
Actin cytoskeleton remodelling drives cell motility, cell to cell contacts, as well as membrane and organelle dynamics. Those cellular activities operate at a particularly high pace in immune cells since these cells migrate through various tissues, interact with multiple cellular partners, ingest microorganisms and secrete effector molecules. The central and multifaceted role of actin cytoskeleton remodelling in sustaining immune cell tasks in humans is highlighted by rare inborn errors of immunity due to mutations in genes encoding proximal and distal actin regulators. In line with the specificity of some of the actin-based processes at work in immune cells, the expression of some of the affected genes, such as WAS, ARPC1B and HEM1 is restricted to the hematopoietic compartment. Exploration of these natural deficiencies highlights the fact that the molecular control of actin remodelling is tuned distinctly in the various subsets of myeloid and lymphoid immune cells and sustains different networks associated with a vast array of specialized tasks. Furthermore, defects in individual actin remodelling proteins are usually associated with partial cellular impairments highlighting the plasticity of actin cytoskeleton remodelling. This review covers the roles of disease-associated actin regulators in promoting the actin-based processes of immune cells. It focuses on the specific molecular function of those regulators across various immune cell subsets and in response to different stimuli. Given the fact that numerous immune-related actin defects have only been characterized recently, we further discuss the challenges lying ahead to decipher the underlying patho-mechanisms. |
Prunier, Guilhèn; Chaves, Beatriz; Lacouture, Claire; Dupré, Loïc Metrics of 2D immunological synapses in human Ŧ cells via high-content confocal cell imaging Journal Article In: Methods Cell Biol, vol. 178, pp. 107–120, 2023, ISSN: 0091-679X. @article{prunier_metrics_2023,
title = {Metrics of 2D immunological synapses in human Ŧ cells via high-content confocal cell imaging},
author = {Prunier, Guilhèn and Chaves, Beatriz and Lacouture, Claire and Dupré, Loïc},
doi = {10.1016/bs.mcb.2022.07.013},
issn = {0091-679X},
year = {2023},
date = {2023-01-01},
journal = {Methods Cell Biol},
volume = {178},
pages = {107--120},
abstract = {Immunological synapses (IS) are the privileged site of complex information transfer between T cells and antigen presenting cells. IS are highly structured in terms of actin and tubulin cytoskeleton organization, receptor and proximal signal patterning, and intracellular organelle polarization. The magnitude and quality of T cell responses upon antigen recognition is dependent on IS molecular organization. For that reason, methods to precisely assess IS parameters are crucial to monitor T cell activation and function in health and disease, but also for T cell centered therapeutic intervention. Confocal and super-resolution microscopy approaches have allowed to characterize the complex structure of the T cell IS. However, those approaches suffer from a low-throughput and low-content format precluding multi-parametric classification of IS across large numbers of samples or stimulatory conditions. Here, we present a protocol of high-content confocal cell imaging in a 384-well plate format adapted to the unbiased analysis of primary T cells forming IS over pre-coated stimulatory molecules. The protocol focuses on the staining of F-actin, pericentrin and granzyme B in CD8+ T cells, but is transposable to other IS molecular markers and lymphocyte subsets. We discuss potential applications offered by the multi-parametric characterization of T cell IS in a high-throughput format.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
Immunological synapses (IS) are the privileged site of complex information transfer between T cells and antigen presenting cells. IS are highly structured in terms of actin and tubulin cytoskeleton organization, receptor and proximal signal patterning, and intracellular organelle polarization. The magnitude and quality of T cell responses upon antigen recognition is dependent on IS molecular organization. For that reason, methods to precisely assess IS parameters are crucial to monitor T cell activation and function in health and disease, but also for T cell centered therapeutic intervention. Confocal and super-resolution microscopy approaches have allowed to characterize the complex structure of the T cell IS. However, those approaches suffer from a low-throughput and low-content format precluding multi-parametric classification of IS across large numbers of samples or stimulatory conditions. Here, we present a protocol of high-content confocal cell imaging in a 384-well plate format adapted to the unbiased analysis of primary T cells forming IS over pre-coated stimulatory molecules. The protocol focuses on the staining of F-actin, pericentrin and granzyme B in CD8+ T cells, but is transposable to other IS molecular markers and lymphocyte subsets. We discuss potential applications offered by the multi-parametric characterization of T cell IS in a high-throughput format. |
Lacouture, Claire; Prunier, Guilhèn; Dupré, Loïc Kinetic measurements of human CD8+ Ŧ cell cytotoxic activity in a 384-well plate format Journal Article In: Methods Cell Biol, vol. 178, pp. 121–133, 2023, ISSN: 0091-679X. @article{lacouture_kinetic_2023,
title = {Kinetic measurements of human CD8+ Ŧ cell cytotoxic activity in a 384-well plate format},
author = {Lacouture, Claire and Prunier, Guilhèn and Dupré, Loïc},
doi = {10.1016/bs.mcb.2022.07.014},
issn = {0091-679X},
year = {2023},
date = {2023-01-01},
journal = {Methods Cell Biol},
volume = {178},
pages = {121--133},
abstract = {The elimination of infected or cancerous cells by CD8+ cytotoxic T lymphocytes (CTL) is a crucial effector mechanism of the immune system. Upon antigen recognition, CTL stop migrating, establish a tight contact with target cells and deliver cytotoxic molecules such as perforin and granzymes that lead to target cell apoptosis. The ability of CTL to control a population of infected cells or a tumor depends on multiple parameters, such as the relative numbers of CTL and target cells, the intrinsic cytotoxic activity of CTL, the intrinsic resistance of target cells and the repertoire of immune checkpoints tuning cytotoxic activity at the CTL:target cell interface. In this context, in vitro assays to precisely measure CTL:target cell interactions and cytotoxic activity over time are required to monitor natural or therapeutic responses. We here present an image-based method that allows recording of positions and survival of CTL and target cells over time in a high-throughput format. The protocol relies on the staining of CTL and target cells with fluorescent dyes and the automated imaging of cells deposited in wells of a 384-well plate with an automated cell imaging device. We discuss potential applications offered by the kinetic assessment of CTL cytotoxic activity in a high-throughput format.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
The elimination of infected or cancerous cells by CD8+ cytotoxic T lymphocytes (CTL) is a crucial effector mechanism of the immune system. Upon antigen recognition, CTL stop migrating, establish a tight contact with target cells and deliver cytotoxic molecules such as perforin and granzymes that lead to target cell apoptosis. The ability of CTL to control a population of infected cells or a tumor depends on multiple parameters, such as the relative numbers of CTL and target cells, the intrinsic cytotoxic activity of CTL, the intrinsic resistance of target cells and the repertoire of immune checkpoints tuning cytotoxic activity at the CTL:target cell interface. In this context, in vitro assays to precisely measure CTL:target cell interactions and cytotoxic activity over time are required to monitor natural or therapeutic responses. We here present an image-based method that allows recording of positions and survival of CTL and target cells over time in a high-throughput format. The protocol relies on the staining of CTL and target cells with fluorescent dyes and the automated imaging of cells deposited in wells of a 384-well plate with an automated cell imaging device. We discuss potential applications offered by the kinetic assessment of CTL cytotoxic activity in a high-throughput format. |
2022
|
Nicoli, Francesco; Cabral-Piccin, Mariela P.; Papagno, Laura; Gallerani, Eleonora; Fusaro, Mathieu; Folcher, Victor; Dubois, Marion; Clave, Emmanuel; Vallet, Hélène; Frere, Justin J.; Gostick, Emma; Llewellyn-Lacey, Sian; Price, David A.; Toubert, Antoine; Dupré, Loïc; Boddaert, Jacques; Caputo, Antonella; Gavioli, Riccardo; Appay, Victor Altered Basal Lipid Metabolism Underlies the Functional Impairment of Naive CD8+ Ŧ Cells in Elderly Humans Journal Article In: J Immunol, vol. 208, no. 3, pp. 562–570, 2022, ISSN: 1550-6606. @article{nicoli_altered_2022b,
title = {Altered Basal Lipid Metabolism Underlies the Functional Impairment of Naive CD8+ Ŧ Cells in Elderly Humans},
author = {Nicoli, Francesco and Cabral-Piccin, Mariela P. and Papagno, Laura and Gallerani, Eleonora and Fusaro, Mathieu and Folcher, Victor and Dubois, Marion and Clave, Emmanuel and Vallet, Hélène and Frere, Justin J. and Gostick, Emma and Llewellyn-Lacey, Sian and Price, David A. and Toubert, Antoine and Dupré, Loïc and Boddaert, Jacques and Caputo, Antonella and Gavioli, Riccardo and Appay, Victor},
doi = {10.4049/jimmunol.2100194},
issn = {1550-6606},
year = {2022},
date = {2022-02-01},
journal = {J Immunol},
volume = {208},
number = {3},
pages = {562--570},
abstract = {Aging is associated with functional deficits in the naive T cell compartment, which compromise the generation of de novo immune responses against previously unencountered Ags. The mechanisms that underlie this phenomenon have nonetheless remained unclear. We found that naive CD8+ T cells in elderly humans were prone to apoptosis and proliferated suboptimally in response to stimulation via the TCR. These abnormalities were associated with dysregulated lipid metabolism under homeostatic conditions and enhanced levels of basal activation. Importantly, reversal of the bioenergetic anomalies with lipid-altering drugs, such as rosiglitazone, almost completely restored the Ag responsiveness of naive CD8+ T cells. Interventions that favor lipid catabolism may therefore find utility as adjunctive therapies in the elderly to promote vaccine-induced immunity against targetable cancers and emerging pathogens, such as seasonal influenza viruses and severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2).},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
Aging is associated with functional deficits in the naive T cell compartment, which compromise the generation of de novo immune responses against previously unencountered Ags. The mechanisms that underlie this phenomenon have nonetheless remained unclear. We found that naive CD8+ T cells in elderly humans were prone to apoptosis and proliferated suboptimally in response to stimulation via the TCR. These abnormalities were associated with dysregulated lipid metabolism under homeostatic conditions and enhanced levels of basal activation. Importantly, reversal of the bioenergetic anomalies with lipid-altering drugs, such as rosiglitazone, almost completely restored the Ag responsiveness of naive CD8+ T cells. Interventions that favor lipid catabolism may therefore find utility as adjunctive therapies in the elderly to promote vaccine-induced immunity against targetable cancers and emerging pathogens, such as seasonal influenza viruses and severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). |
van Steijn, Leonie; Wortel, Inge M. N.; Sire, Clément; Dupré, Loïc; Theraulaz, Guy; Merks, Roeland M. H. Computational modelling of cell motility modes emerging from cell-matrix adhesion dynamics Journal Article In: PLoS Comput Biol, vol. 18, no. 2, pp. e1009156, 2022, ISSN: 1553-7358. @article{van_steijn_computational_2022,
title = {Computational modelling of cell motility modes emerging from cell-matrix adhesion dynamics},
author = {van Steijn, Leonie and Wortel, Inge M. N. and Sire, Clément and Dupré, Loïc and Theraulaz, Guy and Merks, Roeland M. H.},
doi = {10.1371/journal.pcbi.1009156},
issn = {1553-7358},
year = {2022},
date = {2022-01-01},
journal = {PLoS Comput Biol},
volume = {18},
number = {2},
pages = {e1009156},
abstract = {Lymphocytes have been described to perform different motility patterns such as Brownian random walks, persistent random walks, and Lévy walks. Depending on the conditions, such as confinement or the distribution of target cells, either Brownian or Lévy walks lead to more efficient interaction with the targets. The diversity of these motility patterns may be explained by an adaptive response to the surrounding extracellular matrix (ECM). Indeed, depending on the ECM composition, lymphocytes either display a floating motility without attaching to the ECM, or sliding and stepping motility with respectively continuous or discontinuous attachment to the ECM, or pivoting behaviour with sustained attachment to the ECM. Moreover, on the long term, lymphocytes either perform a persistent random walk or a Brownian-like movement depending on the ECM composition. How the ECM affects cell motility is still incompletely understood. Here, we integrate essential mechanistic details of the lymphocyte-matrix adhesions and lymphocyte intrinsic cytoskeletal induced cell propulsion into a Cellular Potts model (CPM). We show that the combination of de novo cell-matrix adhesion formation, adhesion growth and shrinkage, adhesion rupture, and feedback of adhesions onto cell propulsion recapitulates multiple lymphocyte behaviours, for different lymphocyte subsets and various substrates. With an increasing attachment area and increased adhesion strength, the cells' speed and persistence decreases. Additionally, the model predicts random walks with short-term persistent but long-term subdiffusive properties resulting in a pivoting type of motility. For small adhesion areas, the spatial distribution of adhesions emerges as a key factor influencing cell motility. Small adhesions at the front allow for more persistent motility than larger clusters at the back, despite a similar total adhesion area. In conclusion, we present an integrated framework to simulate the effects of ECM proteins on cell-matrix adhesion dynamics. The model reveals a sufficient set of principles explaining the plasticity of lymphocyte motility.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
Lymphocytes have been described to perform different motility patterns such as Brownian random walks, persistent random walks, and Lévy walks. Depending on the conditions, such as confinement or the distribution of target cells, either Brownian or Lévy walks lead to more efficient interaction with the targets. The diversity of these motility patterns may be explained by an adaptive response to the surrounding extracellular matrix (ECM). Indeed, depending on the ECM composition, lymphocytes either display a floating motility without attaching to the ECM, or sliding and stepping motility with respectively continuous or discontinuous attachment to the ECM, or pivoting behaviour with sustained attachment to the ECM. Moreover, on the long term, lymphocytes either perform a persistent random walk or a Brownian-like movement depending on the ECM composition. How the ECM affects cell motility is still incompletely understood. Here, we integrate essential mechanistic details of the lymphocyte-matrix adhesions and lymphocyte intrinsic cytoskeletal induced cell propulsion into a Cellular Potts model (CPM). We show that the combination of de novo cell-matrix adhesion formation, adhesion growth and shrinkage, adhesion rupture, and feedback of adhesions onto cell propulsion recapitulates multiple lymphocyte behaviours, for different lymphocyte subsets and various substrates. With an increasing attachment area and increased adhesion strength, the cells' speed and persistence decreases. Additionally, the model predicts random walks with short-term persistent but long-term subdiffusive properties resulting in a pivoting type of motility. For small adhesion areas, the spatial distribution of adhesions emerges as a key factor influencing cell motility. Small adhesions at the front allow for more persistent motility than larger clusters at the back, despite a similar total adhesion area. In conclusion, we present an integrated framework to simulate the effects of ECM proteins on cell-matrix adhesion dynamics. The model reveals a sufficient set of principles explaining the plasticity of lymphocyte motility. |
Argenty, Jérémy; Rouquié, Nelly; Bories, Cyrielle; Mélique, Suzanne; Duplan-Eche, Valérie; Saoudi, Abdelhadi; Fazilleau, Nicolas; Lesourne, Renaud A selective LIS1 requirement for mitotic spindle assembly discriminates distinct Ŧ-cell division mechanisms within the Ŧ-cell lineage Journal Article In: Elife, vol. 11, pp. e80277, 2022, ISSN: 2050-084X. @article{argenty_selective_2022,
title = {A selective LIS1 requirement for mitotic spindle assembly discriminates distinct Ŧ-cell division mechanisms within the Ŧ-cell lineage},
author = {Argenty, Jérémy and Rouquié, Nelly and Bories, Cyrielle and Mélique, Suzanne and Duplan-Eche, Valérie and Saoudi, Abdelhadi and Fazilleau, Nicolas and Lesourne, Renaud},
doi = {10.7554/eLife.80277},
issn = {2050-084X},
year = {2022},
date = {2022-01-01},
journal = {Elife},
volume = {11},
pages = {e80277},
abstract = {The ability to proliferate is a common feature of most T-cell populations. However, proliferation follows different cell-cycle dynamics and is coupled to different functional outcomes according to T-cell subsets. Whether the mitotic machineries supporting these qualitatively distinct proliferative responses are identical remains unknown. Here, we show that disruption of the microtubule-associated protein LIS1 in mouse models leads to proliferative defects associated with a blockade of T-cell development after β-selection and of peripheral CD4+ T-cell expansion after antigen priming. In contrast, cell divisions in CD8+ T cells occurred independently of LIS1 following T-cell antigen receptor stimulation, although LIS1 was required for proliferation elicited by pharmacological activation. In thymocytes and CD4+ T cells, LIS1 deficiency did not affect signaling events leading to activation but led to an interruption of proliferation after the initial round of division and to p53-induced cell death. Proliferative defects resulted from a mitotic failure, characterized by the presence of extra-centrosomes and the formation of multipolar spindles, causing abnormal chromosomes congression during metaphase and separation during telophase. LIS1 was required to stabilize dynein/dynactin complexes, which promote chromosome attachment to mitotic spindles and ensure centrosome integrity. Together, these results suggest that proliferative responses are supported by distinct mitotic machineries across T-cell subsets.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
The ability to proliferate is a common feature of most T-cell populations. However, proliferation follows different cell-cycle dynamics and is coupled to different functional outcomes according to T-cell subsets. Whether the mitotic machineries supporting these qualitatively distinct proliferative responses are identical remains unknown. Here, we show that disruption of the microtubule-associated protein LIS1 in mouse models leads to proliferative defects associated with a blockade of T-cell development after β-selection and of peripheral CD4+ T-cell expansion after antigen priming. In contrast, cell divisions in CD8+ T cells occurred independently of LIS1 following T-cell antigen receptor stimulation, although LIS1 was required for proliferation elicited by pharmacological activation. In thymocytes and CD4+ T cells, LIS1 deficiency did not affect signaling events leading to activation but led to an interruption of proliferation after the initial round of division and to p53-induced cell death. Proliferative defects resulted from a mitotic failure, characterized by the presence of extra-centrosomes and the formation of multipolar spindles, causing abnormal chromosomes congression during metaphase and separation during telophase. LIS1 was required to stabilize dynein/dynactin complexes, which promote chromosome attachment to mitotic spindles and ensure centrosome integrity. Together, these results suggest that proliferative responses are supported by distinct mitotic machineries across T-cell subsets. |
Mélique, Suzanne; Yang, Cui; Lesourne, Renaud Negative times negative equals positive, THEMIS sets the rule on thymic selection and peripheral Ŧ cell responses Journal Article In: Biomedical Journal, vol. 45, no. 2, pp. 334–346, 2022, ISSN: 2319-4170. @article{melique_negative_2022,
title = {Negative times negative equals positive, THEMIS sets the rule on thymic selection and peripheral Ŧ cell responses},
author = {Mélique, Suzanne and Yang, Cui and Lesourne, Renaud},
url = {https://www.sciencedirect.com/science/article/pii/S2319417022000373},
doi = {10.1016/j.bj.2022.03.008},
issn = {2319-4170},
year = {2022},
date = {2022-01-01},
urldate = {2022-07-20},
journal = {Biomedical Journal},
volume = {45},
number = {2},
pages = {334--346},
abstract = {The activity of T cells is finely controlled by a set of negative regulators of T-cell antigen receptor (TCR)-mediated signaling. However, how those negative regulators are themselves controlled to prevent ineffective TCR-mediated responses remain poorly understood. Thymocyte-expressed molecule involved in selection (THEMIS) has been characterized over a decade ago as an important player of T cell development. Although the molecular function of THEMIS has long remained puzzling and subject to controversies, latest investigations suggest that THEMIS stimulates TCR-mediated signaling by repressing the tyrosine phosphatases SHP-1 and SHP-2 which exert regulatory function on T cell activation. Recent evidences also point to a role for THEMIS in peripheral T cells beyond its role on thymic selection. Here, we present an overview of the past research on THEMIS in the context of T cell development and peripheral T cell function and discuss the possible implication of THEMIS-based mechanisms on TCR-dependent and independent signaling outcomes.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
The activity of T cells is finely controlled by a set of negative regulators of T-cell antigen receptor (TCR)-mediated signaling. However, how those negative regulators are themselves controlled to prevent ineffective TCR-mediated responses remain poorly understood. Thymocyte-expressed molecule involved in selection (THEMIS) has been characterized over a decade ago as an important player of T cell development. Although the molecular function of THEMIS has long remained puzzling and subject to controversies, latest investigations suggest that THEMIS stimulates TCR-mediated signaling by repressing the tyrosine phosphatases SHP-1 and SHP-2 which exert regulatory function on T cell activation. Recent evidences also point to a role for THEMIS in peripheral T cells beyond its role on thymic selection. Here, we present an overview of the past research on THEMIS in the context of T cell development and peripheral T cell function and discuss the possible implication of THEMIS-based mechanisms on TCR-dependent and independent signaling outcomes. |
Yang, Cui; Blaize, Gaëtan; Marrocco, Rémi; Rouquié, Nelly; Bories, Cyrielle; Gador, Mylène; Mélique, Suzanne; Joulia, Emeline; Benamar, Mehdi; Dejean, Anne S.; Daniels-Treffandier, Hélène; Love, Paul E.; Fazilleau, Nicolas; Saoudi, Abdelhadi; Lesourne, Renaud THEMIS enhances the magnitude of normal and neuroinflammatory type 1 immune responses by promoting TCR-independent signals Journal Article In: Science Signaling, vol. 15, no. 742, pp. eabl5343, 2022, (Publisher: American Association for the Advancement of Science). @article{yang_themis_2022,
title = {THEMIS enhances the magnitude of normal and neuroinflammatory type 1 immune responses by promoting TCR-independent signals},
author = {Yang, Cui and Blaize, Gaëtan and Marrocco, Rémi and Rouquié, Nelly and Bories, Cyrielle and Gador, Mylène and Mélique, Suzanne and Joulia, Emeline and Benamar, Mehdi and Dejean, Anne S. and Daniels-Treffandier, Hélène and Love, Paul E. and Fazilleau, Nicolas and Saoudi, Abdelhadi and Lesourne, Renaud},
url = {https://www.science.org/doi/abs/10.1126/scisignal.abl5343},
doi = {10.1126/scisignal.abl5343},
year = {2022},
date = {2022-01-01},
urldate = {2022-01-01},
journal = {Science Signaling},
volume = {15},
number = {742},
pages = {eabl5343},
note = {Publisher: American Association for the Advancement of Science},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
|
Magnani, A.; Semeraro, M.; Adam, F.; Booth, C.; Dupré, L.; Morris, E. C.; Gabrion, A.; Roudaut, C.; Borgel, D.; Toubert, A.; Clave, E.; Abdo, C.; Gorochov, G.; Petermann, R.; Guiot, M.; Miyara, M.; Moshous, D.; Magrin, E.; Denis, A.; Suarez, F.; Lagresle, C.; Roche, A. M.; Everett, J.; Trinquand, A.; Guisset, M.; Bayford, J. Xu; Hacein-Bey-Abina, S.; Kauskot, A.; Elfeky, R.; Rivat, C.; Abbas, S.; Gaspar, H. B.; Macintyre, E.; Picard, C.; Bushman, F. D.; Galy, A.; Fischer, A.; Six, E.; Thrasher, A. J.; Cavazzana, M. Long-term safety and efficacy of lentiviral hematopoietic stem/progenitor cell gene therapy for Wiskott–Aldrich syndrome Journal Article In: Nat Med, pp. 1–10, 2022, ISSN: 1546-170X, (Bandiera_abtest: a
Cc_license_type: cc_by
Cg_type: Nature Research Journals
Primary_atype: Research
Publisher: Nature Publishing Group
Subject_term: Primary immunodeficiency disorders;Targeted gene repair
Subject_term_id: primary-immunodeficiency-disorders;targeted-gene-repair). @article{magnani_long-term_2022,
title = {Long-term safety and efficacy of lentiviral hematopoietic stem/progenitor cell gene therapy for Wiskott–Aldrich syndrome},
author = {Magnani, A. and Semeraro, M. and Adam, F. and Booth, C. and Dupré, L. and Morris, E. C. and Gabrion, A. and Roudaut, C. and Borgel, D. and Toubert, A. and Clave, E. and Abdo, C. and Gorochov, G. and Petermann, R. and Guiot, M. and Miyara, M. and Moshous, D. and Magrin, E. and Denis, A. and Suarez, F. and Lagresle, C. and Roche, A. M. and Everett, J. and Trinquand, A. and Guisset, M. and Bayford, J. Xu and Hacein-Bey-Abina, S. and Kauskot, A. and Elfeky, R. and Rivat, C. and Abbas, S. and Gaspar, H. B. and Macintyre, E. and Picard, C. and Bushman, F. D. and Galy, A. and Fischer, A. and Six, E. and Thrasher, A. J. and Cavazzana, M.},
url = {https://www.nature.com/articles/s41591-021-01641-x},
doi = {10.1038/s41591-021-01641-x},
issn = {1546-170X},
year = {2022},
date = {2022-01-01},
urldate = {2022-01-26},
journal = {Nat Med},
pages = {1--10},
abstract = {Patients with Wiskott–Aldrich syndrome (WAS) lacking a human leukocyte antigen-matched donor may benefit from gene therapy through the provision of gene-corrected, autologous hematopoietic stem/progenitor cells. Here, we present comprehensive, long-term follow-up results (median follow-up, 7.6 years) (phase I/II trial no. NCT02333760) for eight patients with WAS having undergone phase I/II lentiviral vector-based gene therapy trials (nos. NCT01347346and NCT01347242), with a focus on thrombocytopenia and autoimmunity. Primary outcomes of the long-term study were to establish clinical and biological safety, efficacy and tolerability by evaluating the incidence and type of serious adverse events and clinical status and biological parameters including lentiviral genomic integration sites in different cell subpopulations from 3 years to 15 years after gene therapy. Secondary outcomes included monitoring the need for additional treatment and T cell repertoire diversity. An interim analysis shows that the study meets the primary outcome criteria tested given that the gene-corrected cells engrafted stably, and no serious treatment-associated adverse events occurred. Overall, severe infections and eczema resolved. Autoimmune disorders and bleeding episodes were significantly less frequent, despite only partial correction of the platelet compartment. The results suggest that lentiviral gene therapy provides sustained clinical benefits for patients with WAS.},
note = {Bandiera_abtest: a
Cc_license_type: cc_by
Cg_type: Nature Research Journals
Primary_atype: Research
Publisher: Nature Publishing Group
Subject_term: Primary immunodeficiency disorders;Targeted gene repair
Subject_term_id: primary-immunodeficiency-disorders;targeted-gene-repair},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
Patients with Wiskott–Aldrich syndrome (WAS) lacking a human leukocyte antigen-matched donor may benefit from gene therapy through the provision of gene-corrected, autologous hematopoietic stem/progenitor cells. Here, we present comprehensive, long-term follow-up results (median follow-up, 7.6 years) (phase I/II trial no. NCT02333760) for eight patients with WAS having undergone phase I/II lentiviral vector-based gene therapy trials (nos. NCT01347346and NCT01347242), with a focus on thrombocytopenia and autoimmunity. Primary outcomes of the long-term study were to establish clinical and biological safety, efficacy and tolerability by evaluating the incidence and type of serious adverse events and clinical status and biological parameters including lentiviral genomic integration sites in different cell subpopulations from 3 years to 15 years after gene therapy. Secondary outcomes included monitoring the need for additional treatment and T cell repertoire diversity. An interim analysis shows that the study meets the primary outcome criteria tested given that the gene-corrected cells engrafted stably, and no serious treatment-associated adverse events occurred. Overall, severe infections and eczema resolved. Autoimmune disorders and bleeding episodes were significantly less frequent, despite only partial correction of the platelet compartment. The results suggest that lentiviral gene therapy provides sustained clinical benefits for patients with WAS. |
2021
|
Vulliard, Loan; Hancock, Joel; Kamnev, Anton; Fell, Christopher W.; Ferreira da Silva, Joana; Loizou, Joanna I.; Nagy, Vanja; Dupré, Loïc; Menche, Jörg BioProfiling.jl: Profiling biological perturbations with high-content imaging in single cells and heterogeneous populations Journal Article In: Bioinformatics, pp. btab853, 2021, ISSN: 1367-4811. @article{vulliard_bioprofilingjl_2021b,
title = {BioProfiling.jl: Profiling biological perturbations with high-content imaging in single cells and heterogeneous populations},
author = {Vulliard, Loan and Hancock, Joel and Kamnev, Anton and Fell, Christopher W. and Ferreira da Silva, Joana and Loizou, Joanna I. and Nagy, Vanja and Dupré, Loïc and Menche, Jörg},
doi = {10.1093/bioinformatics/btab853},
issn = {1367-4811},
year = {2021},
date = {2021-12-01},
journal = {Bioinformatics},
pages = {btab853},
abstract = {MOTIVATION: High-content imaging screens provide a cost-effective and scalable way to assess cell states across diverse experimental conditions. The analysis of the acquired microscopy images involves assembling and curating raw cellular measurements into morphological profiles suitable for testing biological hypotheses. Despite being a critical step, general-purpose and adaptable tools for morphological profiling are lacking and no solution is available for the high-performance Julia programming language.
RESULTS: Here, we introduce BioProfiling.jl, an efficient end-to-end solution for compiling and filtering informative morphological profiles in Julia. The package contains all the necessary data structures to curate morphological measurements and helper functions to transform, normalize and visualize profiles. Robust statistical distances and permutation tests enable quantification of the significance of the observed changes despite the high fraction of outliers inherent to high-content screens. This package also simplifies visual artifact diagnostics, thus streamlining a bottleneck of morphological analyses. We showcase the features of the package by analyzing a chemical imaging screen, in which the morphological profiles prove to be informative about the compounds' mechanisms of action and can be conveniently integrated with the network localization of molecular targets.
AVAILABILITY: The Julia package is available on GitHub: https://github.com/menchelab/BioProfiling.jlWe also provide Jupyter notebooks reproducing our analyses: https://github.com/menchelab/BioProfilingNotebooks.
SUPPLEMENTARY INFORMATION: Supplementary data are available at Bioinformatics online.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
MOTIVATION: High-content imaging screens provide a cost-effective and scalable way to assess cell states across diverse experimental conditions. The analysis of the acquired microscopy images involves assembling and curating raw cellular measurements into morphological profiles suitable for testing biological hypotheses. Despite being a critical step, general-purpose and adaptable tools for morphological profiling are lacking and no solution is available for the high-performance Julia programming language.
RESULTS: Here, we introduce BioProfiling.jl, an efficient end-to-end solution for compiling and filtering informative morphological profiles in Julia. The package contains all the necessary data structures to curate morphological measurements and helper functions to transform, normalize and visualize profiles. Robust statistical distances and permutation tests enable quantification of the significance of the observed changes despite the high fraction of outliers inherent to high-content screens. This package also simplifies visual artifact diagnostics, thus streamlining a bottleneck of morphological analyses. We showcase the features of the package by analyzing a chemical imaging screen, in which the morphological profiles prove to be informative about the compounds' mechanisms of action and can be conveniently integrated with the network localization of molecular targets.
AVAILABILITY: The Julia package is available on GitHub: https://github.com/menchelab/BioProfiling.jlWe also provide Jupyter notebooks reproducing our analyses: https://github.com/menchelab/BioProfilingNotebooks.
SUPPLEMENTARY INFORMATION: Supplementary data are available at Bioinformatics online. |
Kamnev, Anton; Lacouture, Claire; Fusaro, Mathieu; Dupré, Loïc Molecular Tuning of Actin Dynamics in Leukocyte Migration as Revealed by Immune-Related Actinopathies Journal Article In: Front Immunol, vol. 12, pp. 750537, 2021, ISSN: 1664-3224. @article{kamnev_molecular_2021b,
title = {Molecular Tuning of Actin Dynamics in Leukocyte Migration as Revealed by Immune-Related Actinopathies},
author = {Kamnev, Anton and Lacouture, Claire and Fusaro, Mathieu and Dupré, Loïc},
doi = {10.3389/fimmu.2021.750537},
issn = {1664-3224},
year = {2021},
date = {2021-01-01},
journal = {Front Immunol},
volume = {12},
pages = {750537},
abstract = {Motility is a crucial activity of immune cells allowing them to patrol tissues as they differentiate, sample or exchange information, and execute their effector functions. Although all immune cells are highly migratory, each subset is endowed with very distinct motility patterns in accordance with functional specification. Furthermore individual immune cell subsets adapt their motility behaviour to the surrounding tissue environment. This review focuses on how the generation and adaptation of diversified motility patterns in immune cells is sustained by actin cytoskeleton dynamics. In particular, we review the knowledge gained through the study of inborn errors of immunity (IEI) related to actin defects. Such pathologies are unique models that help us to uncover the contribution of individual actin regulators to the migration of immune cells in the context of their development and function.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
Motility is a crucial activity of immune cells allowing them to patrol tissues as they differentiate, sample or exchange information, and execute their effector functions. Although all immune cells are highly migratory, each subset is endowed with very distinct motility patterns in accordance with functional specification. Furthermore individual immune cell subsets adapt their motility behaviour to the surrounding tissue environment. This review focuses on how the generation and adaptation of diversified motility patterns in immune cells is sustained by actin cytoskeleton dynamics. In particular, we review the knowledge gained through the study of inborn errors of immunity (IEI) related to actin defects. Such pathologies are unique models that help us to uncover the contribution of individual actin regulators to the migration of immune cells in the context of their development and function. |
Dupré, Loïc; Boztug, Kaan; Pfajfer, Laurène Actin Dynamics at the Ŧ Cell Synapse as Revealed by Immune-Related Actinopathies Journal Article In: Front Cell Dev Biol, vol. 9, pp. 665519, 2021, ISSN: 2296-634X. @article{dupre_actin_2021b,
title = {Actin Dynamics at the Ŧ Cell Synapse as Revealed by Immune-Related Actinopathies},
author = {Dupré, Loïc and Boztug, Kaan and Pfajfer, Laurène},
doi = {10.3389/fcell.2021.665519},
issn = {2296-634X},
year = {2021},
date = {2021-01-01},
journal = {Front Cell Dev Biol},
volume = {9},
pages = {665519},
abstract = {The actin cytoskeleton is composed of dynamic filament networks that build adaptable local architectures to sustain nearly all cellular activities in response to a myriad of stimuli. Although the function of numerous players that tune actin remodeling is known, the coordinated molecular orchestration of the actin cytoskeleton to guide cellular decisions is still ill defined. T lymphocytes provide a prototypical example of how a complex program of actin cytoskeleton remodeling sustains the spatio-temporal control of key cellular activities, namely antigen scanning and sensing, as well as polarized delivery of effector molecules, via the immunological synapse. We here review the unique knowledge on actin dynamics at the T lymphocyte synapse gained through the study of primary immunodeficiences caused by mutations in genes encoding actin regulatory proteins. Beyond the specific roles of individual actin remodelers, we further develop the view that these operate in a coordinated manner and are an integral part of multiple signaling pathways in T lymphocytes.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
The actin cytoskeleton is composed of dynamic filament networks that build adaptable local architectures to sustain nearly all cellular activities in response to a myriad of stimuli. Although the function of numerous players that tune actin remodeling is known, the coordinated molecular orchestration of the actin cytoskeleton to guide cellular decisions is still ill defined. T lymphocytes provide a prototypical example of how a complex program of actin cytoskeleton remodeling sustains the spatio-temporal control of key cellular activities, namely antigen scanning and sensing, as well as polarized delivery of effector molecules, via the immunological synapse. We here review the unique knowledge on actin dynamics at the T lymphocyte synapse gained through the study of primary immunodeficiences caused by mutations in genes encoding actin regulatory proteins. Beyond the specific roles of individual actin remodelers, we further develop the view that these operate in a coordinated manner and are an integral part of multiple signaling pathways in T lymphocytes. |
Hamminger, Patricia; Marchetti, Luca; Preglej, Teresa; Platzer, René; Zhu, Ci; Kamnev, Anton; Rica, Ramona; Stolz, Valentina; Sandner, Lisa; Alteneder, Marlis; Kaba, Elisa; Waltenberger, Darina; Huppa, Johannes B.; Trauner, Michael; Bock, Christoph; Lyck, Ruth; Bauer, Jan; Dupré, Loïc; Seiser, Christian; Boucheron, Nicole; Engelhardt, Britta; Ellmeier, Wilfried Histone deacetylase 1 controls CD4+ Ŧ cell trafficking in autoinflammatory diseases Journal Article In: J Autoimmun, vol. 119, pp. 102610, 2021, ISSN: 1095-9157. @article{hamminger_histone_2021b,
title = {Histone deacetylase 1 controls CD4+ Ŧ cell trafficking in autoinflammatory diseases},
author = {Hamminger, Patricia and Marchetti, Luca and Preglej, Teresa and Platzer, René and Zhu, Ci and Kamnev, Anton and Rica, Ramona and Stolz, Valentina and Sandner, Lisa and Alteneder, Marlis and Kaba, Elisa and Waltenberger, Darina and Huppa, Johannes B. and Trauner, Michael and Bock, Christoph and Lyck, Ruth and Bauer, Jan and Dupré, Loïc and Seiser, Christian and Boucheron, Nicole and Engelhardt, Britta and Ellmeier, Wilfried},
doi = {10.1016/j.jaut.2021.102610},
issn = {1095-9157},
year = {2021},
date = {2021-01-01},
journal = {J Autoimmun},
volume = {119},
pages = {102610},
abstract = {CD4+ T cell trafficking is a fundamental property of adaptive immunity. In this study, we uncover a novel role for histone deacetylase 1 (HDAC1) in controlling effector CD4+ T cell migration, thereby providing mechanistic insight into why a T cell-specific deletion of HDAC1 protects against experimental autoimmune encephalomyelitis (EAE). HDAC1-deficient CD4+ T cells downregulated genes associated with leukocyte extravasation. In vitro, HDAC1-deficient CD4+ T cells displayed aberrant morphology and migration on surfaces coated with integrin LFA-1 ligand ICAM-1 and showed an impaired ability to arrest on and to migrate across a monolayer of primary mouse brain microvascular endothelial cells under physiological flow. Moreover, HDAC1 deficiency reduced homing of CD4+ T cells into the intestinal epithelium and lamina propria preventing weight-loss, crypt damage and intestinal inflammation in adoptive CD4+ T cell transfer colitis. This correlated with reduced expression levels of LFA-1 integrin chains CD11a and CD18 as well as of selectin ligands CD43, CD44 and CD162 on transferred circulating HDAC1-deficient CD4+ T cells. Our data reveal that HDAC1 controls T cell-mediated autoimmunity via the regulation of CD4+ T cell trafficking into the CNS and intestinal tissues.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
CD4+ T cell trafficking is a fundamental property of adaptive immunity. In this study, we uncover a novel role for histone deacetylase 1 (HDAC1) in controlling effector CD4+ T cell migration, thereby providing mechanistic insight into why a T cell-specific deletion of HDAC1 protects against experimental autoimmune encephalomyelitis (EAE). HDAC1-deficient CD4+ T cells downregulated genes associated with leukocyte extravasation. In vitro, HDAC1-deficient CD4+ T cells displayed aberrant morphology and migration on surfaces coated with integrin LFA-1 ligand ICAM-1 and showed an impaired ability to arrest on and to migrate across a monolayer of primary mouse brain microvascular endothelial cells under physiological flow. Moreover, HDAC1 deficiency reduced homing of CD4+ T cells into the intestinal epithelium and lamina propria preventing weight-loss, crypt damage and intestinal inflammation in adoptive CD4+ T cell transfer colitis. This correlated with reduced expression levels of LFA-1 integrin chains CD11a and CD18 as well as of selectin ligands CD43, CD44 and CD162 on transferred circulating HDAC1-deficient CD4+ T cells. Our data reveal that HDAC1 controls T cell-mediated autoimmunity via the regulation of CD4+ T cell trafficking into the CNS and intestinal tissues. |
Kalinichenko, Artem; Perinetti Casoni, Giovanna; Dupré, Loïc; Trotta, Luca; Huemer, Jakob; Galgano, Donatella; German, Yolla; Haladik, Ben; Pazmandi, Julia; Thian, Marini; Yüce Petronczki, Özlem; Chiang, Samuel C.; Taskinen, Mervi; Hekkala, Anne; Kauppila, Saila; Lindgren, Outi; Tapiainen, Terhi; Kraakman, Michael J.; Vettenranta, Kim; Lomakin, Alexis J.; Saarela, Janna; Seppänen, Mikko R. J.; Bryceson, Yenan T.; Boztug, Kaan RhoG deficiency abrogates cytotoxicity of human lymphocytes and causes hemophagocytic lymphohistiocytosis Journal Article In: Blood, vol. 137, no. 15, pp. 2033–2045, 2021, ISSN: 1528-0020. @article{kalinichenko_rhog_2021b,
title = {RhoG deficiency abrogates cytotoxicity of human lymphocytes and causes hemophagocytic lymphohistiocytosis},
author = {Kalinichenko, Artem and Perinetti Casoni, Giovanna and Dupré, Loïc and Trotta, Luca and Huemer, Jakob and Galgano, Donatella and German, Yolla and Haladik, Ben and Pazmandi, Julia and Thian, Marini and Yüce Petronczki, Özlem and Chiang, Samuel C. and Taskinen, Mervi and Hekkala, Anne and Kauppila, Saila and Lindgren, Outi and Tapiainen, Terhi and Kraakman, Michael J. and Vettenranta, Kim and Lomakin, Alexis J. and Saarela, Janna and Seppänen, Mikko R. J. and Bryceson, Yenan T. and Boztug, Kaan},
doi = {10.1182/blood.2020008738},
issn = {1528-0020},
year = {2021},
date = {2021-01-01},
journal = {Blood},
volume = {137},
number = {15},
pages = {2033--2045},
abstract = {Exocytosis of cytotoxic granules (CG) by lymphocytes is required for the elimination of infected and malignant cells. Impairments in this process underly a group of diseases with dramatic hyperferritinemic inflammation termed hemophagocytic lymphohistiocytosis (HLH). Although genetic and functional studies of HLH have identified proteins controlling distinct steps of CG exocytosis, the molecular mechanisms that spatiotemporally coordinate CG release remain partially elusive. We studied a patient exhibiting characteristic clinical features of HLH associated with markedly impaired cytotoxic T lymphocyte (CTL) and natural killer (NK) cell exocytosis functions, who beared biallelic deleterious mutations in the gene encoding the small GTPase RhoG. Experimental ablation of RHOG in a model cell line and primary CTLs from healthy individuals uncovered a hitherto unappreciated role of RhoG in retaining CGs in the vicinity of the plasma membrane (PM), a fundamental prerequisite for CG exocytotic release. We discovered that RhoG engages in a protein-protein interaction with Munc13-4, an exocytosis protein essential for CG fusion with the PM. We show that this interaction is critical for docking of Munc13-4+ CGs to the PM and subsequent membrane fusion and release of CG content. Thus, our study illuminates RhoG as a novel essential regulator of human lymphocyte cytotoxicity and provides the molecular pathomechanism behind the identified here and previously unreported genetically determined form of HLH.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
Exocytosis of cytotoxic granules (CG) by lymphocytes is required for the elimination of infected and malignant cells. Impairments in this process underly a group of diseases with dramatic hyperferritinemic inflammation termed hemophagocytic lymphohistiocytosis (HLH). Although genetic and functional studies of HLH have identified proteins controlling distinct steps of CG exocytosis, the molecular mechanisms that spatiotemporally coordinate CG release remain partially elusive. We studied a patient exhibiting characteristic clinical features of HLH associated with markedly impaired cytotoxic T lymphocyte (CTL) and natural killer (NK) cell exocytosis functions, who beared biallelic deleterious mutations in the gene encoding the small GTPase RhoG. Experimental ablation of RHOG in a model cell line and primary CTLs from healthy individuals uncovered a hitherto unappreciated role of RhoG in retaining CGs in the vicinity of the plasma membrane (PM), a fundamental prerequisite for CG exocytotic release. We discovered that RhoG engages in a protein-protein interaction with Munc13-4, an exocytosis protein essential for CG fusion with the PM. We show that this interaction is critical for docking of Munc13-4+ CGs to the PM and subsequent membrane fusion and release of CG content. Thus, our study illuminates RhoG as a novel essential regulator of human lymphocyte cytotoxicity and provides the molecular pathomechanism behind the identified here and previously unreported genetically determined form of HLH. |
German, Yolla; Vulliard, Loan; Kamnev, Anton; Pfajfer, Laurène; Huemer, Jakob; Mautner, Anna-Katharina; Rubio, Aude; Kalinichenko, Artem; Boztug, Kaan; Ferrand, Audrey; Menche, Jörg; Dupré, Loïc Morphological profiling of human Ŧ and NK lymphocytes by high-content cell imaging Journal Article In: Cell Rep, vol. 36, no. 1, pp. 109318, 2021, ISSN: 2211-1247. @article{german_morphological_2021b,
title = {Morphological profiling of human Ŧ and NK lymphocytes by high-content cell imaging},
author = {German, Yolla and Vulliard, Loan and Kamnev, Anton and Pfajfer, Laurène and Huemer, Jakob and Mautner, Anna-Katharina and Rubio, Aude and Kalinichenko, Artem and Boztug, Kaan and Ferrand, Audrey and Menche, Jörg and Dupré, Loïc},
doi = {10.1016/j.celrep.2021.109318},
issn = {2211-1247},
year = {2021},
date = {2021-01-01},
journal = {Cell Rep},
volume = {36},
number = {1},
pages = {109318},
abstract = {The immunological synapse is a complex structure that decodes stimulatory signals into adapted lymphocyte responses. It is a unique window to monitor lymphocyte activity because of development of systematic quantitative approaches. Here we demonstrate the applicability of high-content imaging to human T and natural killer (NK) cells and develop a pipeline for unbiased analysis of high-definition morphological profiles. Our approach reveals how distinct facets of actin cytoskeleton remodeling shape immunological synapse architecture and affect lytic granule positioning. Morphological profiling of CD8+ T cells from immunodeficient individuals allows discrimination of the roles of the ARP2/3 subunit ARPC1B and the ARP2/3 activator Wiskott-Aldrich syndrome protein (WASP) in immunological synapse assembly. Single-cell analysis further identifies uncoupling of lytic granules and F-actin radial distribution in ARPC1B-deficient lymphocytes. Our study provides a foundation for development of morphological profiling as a scalable approach to monitor primary lymphocyte responsiveness and to identify complex aspects of lymphocyte micro-architecture.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
The immunological synapse is a complex structure that decodes stimulatory signals into adapted lymphocyte responses. It is a unique window to monitor lymphocyte activity because of development of systematic quantitative approaches. Here we demonstrate the applicability of high-content imaging to human T and natural killer (NK) cells and develop a pipeline for unbiased analysis of high-definition morphological profiles. Our approach reveals how distinct facets of actin cytoskeleton remodeling shape immunological synapse architecture and affect lytic granule positioning. Morphological profiling of CD8+ T cells from immunodeficient individuals allows discrimination of the roles of the ARP2/3 subunit ARPC1B and the ARP2/3 activator Wiskott-Aldrich syndrome protein (WASP) in immunological synapse assembly. Single-cell analysis further identifies uncoupling of lytic granules and F-actin radial distribution in ARPC1B-deficient lymphocytes. Our study provides a foundation for development of morphological profiling as a scalable approach to monitor primary lymphocyte responsiveness and to identify complex aspects of lymphocyte micro-architecture. |
Hanaei, Sara; Mohebi, Farnam; Moradi-Lakeh, Maziar; Jabbari, Parnian; Mehta, Surinder Kumar; Kryvenko, Liudmyla S.; Luongo, Livio; Dupré, Loďc; Rezaei, Nima The Epidemiologic Aspects of COVID-19 Outbreak: Spreading Beyond Expectations Journal Article In: Adv Exp Med Biol, vol. 1318, pp. 61–79, 2021, ISSN: 0065-2598. @article{hanaei_epidemiologic_2021b,
title = {The Epidemiologic Aspects of COVID-19 Outbreak: Spreading Beyond Expectations},
author = {Hanaei, Sara and Mohebi, Farnam and Moradi-Lakeh, Maziar and Jabbari, Parnian and Mehta, Surinder Kumar and Kryvenko, Liudmyla S. and Luongo, Livio and Dupré, Loďc and Rezaei, Nima},
doi = {10.1007/978-3-030-63761-3_4},
issn = {0065-2598},
year = {2021},
date = {2021-01-01},
journal = {Adv Exp Med Biol},
volume = {1318},
pages = {61--79},
abstract = {The coronavirus disease 2019 (COVID-19) outbreak started in late 2019 in Wuhan, Hubei Province of China, and quickly spread to the surrounding regions and neighboring countries. A novel coronavirus, the so-called severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), was found to be responsible for this outbreak potentially originating from pangolins. In China, the outbreak lasted for 1 month until it seemed to be controlled after affecting over 81,000 individuals and causing deaths in over 4200 patients. Subsequently, and after affecting over 118,000 individuals and causing over 4200 deaths, the condition was officially announced as a pandemic by the World Health Organization (WHO). In the meantime, the epidemic curve took a downtrend in China, the original epicenter of the pandemic, but started to rise in other countries with a steep slope. Among over 215 affected countries, the USA, European countries (Italy, Germany, Spain, France, the UK), Iran, and South Korea had the highest frequencies in the matters of infected patients and deaths. Importantly, different countries took different policies when encountered with an outbreak, especially in the matter of accuracy of the report and timing of the action. A part of the delays in reporting was expected, including the lag in the chain of reporting, the shortcomings of tests, missed patients, and inadequate testing facilities. However, there were also political and nontechnical reasons that caused the reporting to be inaccurate. Surveillance seems to be less of a reason for the observed in poor management, and it mostly originated from human decision-making failures and political issues. Besides, the culture of populations and their trust in their governments played an important role on how they reacted to the COVID-19 pandemic and acquired policies. Finally, the characteristics of the world today indicate the danger of probable upcoming outbreaks, and policymakers should utilize the existing opportunities, particularly the advancements in technology and media, to prevent or adequately manage them.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
The coronavirus disease 2019 (COVID-19) outbreak started in late 2019 in Wuhan, Hubei Province of China, and quickly spread to the surrounding regions and neighboring countries. A novel coronavirus, the so-called severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), was found to be responsible for this outbreak potentially originating from pangolins. In China, the outbreak lasted for 1 month until it seemed to be controlled after affecting over 81,000 individuals and causing deaths in over 4200 patients. Subsequently, and after affecting over 118,000 individuals and causing over 4200 deaths, the condition was officially announced as a pandemic by the World Health Organization (WHO). In the meantime, the epidemic curve took a downtrend in China, the original epicenter of the pandemic, but started to rise in other countries with a steep slope. Among over 215 affected countries, the USA, European countries (Italy, Germany, Spain, France, the UK), Iran, and South Korea had the highest frequencies in the matters of infected patients and deaths. Importantly, different countries took different policies when encountered with an outbreak, especially in the matter of accuracy of the report and timing of the action. A part of the delays in reporting was expected, including the lag in the chain of reporting, the shortcomings of tests, missed patients, and inadequate testing facilities. However, there were also political and nontechnical reasons that caused the reporting to be inaccurate. Surveillance seems to be less of a reason for the observed in poor management, and it mostly originated from human decision-making failures and political issues. Besides, the culture of populations and their trust in their governments played an important role on how they reacted to the COVID-19 pandemic and acquired policies. Finally, the characteristics of the world today indicate the danger of probable upcoming outbreaks, and policymakers should utilize the existing opportunities, particularly the advancements in technology and media, to prevent or adequately manage them. |
2020
|
Donnadieu, Emmanuel; Dupré, Loïc; Pinho, Lia Gonçalves; Cotta-de-Almeida, Vinicius Surmounting the obstacles that impede effective CAR Ŧ cell trafficking to solid tumors Journal Article In: Journal of Leukocyte Biology, 2020, ISSN: 1938-3673. @article{donnadieu_surmounting_2020,
title = {Surmounting the obstacles that impede effective CAR Ŧ cell trafficking to solid tumors},
author = {Donnadieu, Emmanuel and Dupré, Loïc and Pinho, Lia Gonçalves and Cotta-de-Almeida, Vinicius},
doi = {10.1002/JLB.1MR0520-746R},
issn = {1938-3673},
year = {2020},
date = {2020-07-01},
journal = {Journal of Leukocyte Biology},
abstract = {Innovative immunotherapies based on immune checkpoint targeting antibodies and engineered T cells are transforming the way we approach cancer treatment. However, although these T cell centered strategies result in marked and durable responses in patients across many different tumor types, they provide therapeutic efficacy only in a proportion of patients. A major challenge of immuno-oncology is thereby to identify mechanisms responsible for resistance to cancer immunotherapy in order to overcome them via adapted strategies that will ultimately improve intrinsic efficacy and response rates. Here, we focus on the barriers that restrain the trafficking of chimeric antigen receptor (CAR)-expressing T cells to solid tumors. Upon infusion, CAR T cells need to home into malignant sites, navigate within complex tumor environments, form productive interactions with cancer cells, deliver their cytotoxic activities, and finally persist. We review the accumulating evidence that the microenvironment of solid tumors contains multiple obstacles that hinder CAR T cells in the dynamic steps underlying their trafficking. We focus on how these hurdles may in part account for the failure of CAR T cell clinical trials in human carcinomas. Given the engineered nature of CAR T cells and possibilities to modify the tumor environment, there are ample opportunities to augment CAR T cell ability to efficiently find and combat tumors. We present some of these strategies, which represent a dynamic field of research with high potential for clinical applicability.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
Innovative immunotherapies based on immune checkpoint targeting antibodies and engineered T cells are transforming the way we approach cancer treatment. However, although these T cell centered strategies result in marked and durable responses in patients across many different tumor types, they provide therapeutic efficacy only in a proportion of patients. A major challenge of immuno-oncology is thereby to identify mechanisms responsible for resistance to cancer immunotherapy in order to overcome them via adapted strategies that will ultimately improve intrinsic efficacy and response rates. Here, we focus on the barriers that restrain the trafficking of chimeric antigen receptor (CAR)-expressing T cells to solid tumors. Upon infusion, CAR T cells need to home into malignant sites, navigate within complex tumor environments, form productive interactions with cancer cells, deliver their cytotoxic activities, and finally persist. We review the accumulating evidence that the microenvironment of solid tumors contains multiple obstacles that hinder CAR T cells in the dynamic steps underlying their trafficking. We focus on how these hurdles may in part account for the failure of CAR T cell clinical trials in human carcinomas. Given the engineered nature of CAR T cells and possibilities to modify the tumor environment, there are ample opportunities to augment CAR T cell ability to efficiently find and combat tumors. We present some of these strategies, which represent a dynamic field of research with high potential for clinical applicability. |
Lamsoul, Isabelle; Dupré, Loïc; Lutz, Pierre G. Molecular Tuning of Filamin A Activities in the Context of Adhesion and Migration Journal Article In: Front Cell Dev Biol, vol. 8, pp. 591323, 2020, ISSN: 2296-634X. @article{lamsoul_molecular_2020,
title = {Molecular Tuning of Filamin A Activities in the Context of Adhesion and Migration},
author = {Lamsoul, Isabelle and Dupré, Loïc and Lutz, Pierre G.},
doi = {10.3389/fcell.2020.591323},
issn = {2296-634X},
year = {2020},
date = {2020-01-01},
journal = {Front Cell Dev Biol},
volume = {8},
pages = {591323},
abstract = {The dynamic organization of actin cytoskeleton meshworks relies on multiple actin-binding proteins endowed with distinct actin-remodeling activities. Filamin A is a large multi-domain scaffolding protein that cross-links actin filaments with orthogonal orientation in response to various stimuli. As such it plays key roles in the modulation of cell shape, cell motility, and differentiation throughout development and adult life. The essentiality and complexity of Filamin A is highlighted by mutations that lead to a variety of severe human disorders affecting multiple organs. One of the most conserved activity of Filamin A is to bridge the actin cytoskeleton to integrins, thereby maintaining the later in an inactive state. We here review the numerous mechanisms cells have developed to adjust Filamin A content and activity and focus on the function of Filamin A as a gatekeeper to integrin activation and associated adhesion and motility.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
The dynamic organization of actin cytoskeleton meshworks relies on multiple actin-binding proteins endowed with distinct actin-remodeling activities. Filamin A is a large multi-domain scaffolding protein that cross-links actin filaments with orthogonal orientation in response to various stimuli. As such it plays key roles in the modulation of cell shape, cell motility, and differentiation throughout development and adult life. The essentiality and complexity of Filamin A is highlighted by mutations that lead to a variety of severe human disorders affecting multiple organs. One of the most conserved activity of Filamin A is to bridge the actin cytoskeleton to integrins, thereby maintaining the later in an inactive state. We here review the numerous mechanisms cells have developed to adjust Filamin A content and activity and focus on the function of Filamin A as a gatekeeper to integrin activation and associated adhesion and motility. |
Weulersse, Marianne; Asrir, Assia; Pichler, Andrea C.; Lemaitre, Lea; Braun, Matthias; Carrié, Nadège; Joubert, Marie-Véronique; Le Moine, Marie; Do Souto, Laura; Gaud, Guillaume; Das, Indrajit; Brauns, Elisa; Scarlata, Clara M.; Morandi, Elena; Sundarrajan, Ashmitha; Cuisinier, Marine; Buisson, Laure; Maheo, Sabrina; Kassem, Sahar; Agesta, Arantxa; Pérès, Michaël; Verhoeyen, Els; Martinez, Alejandra; Mazieres, Julien; Dupré, Loïc; Gossye, Thomas; Pancaldi, Vera; Guillerey, Camille; Ayyoub, Maha; Dejean, Anne S.; Saoudi, Abdelhadi; Goriely, Stanislas; Avet-Loiseau, Hervé; Bald, Tobias; Smyth, Mark J.; Martinet, Ludovic Eomes-Dependent Loss of the Co-activating Receptor CD226 Restrains CD8+ Ŧ Cell Anti-tumor Functions and Limits the Efficacy of Cancer Immunotherapy Journal Article In: Immunity, vol. 53, no. 4, pp. 824–839.e10, 2020, ISSN: 1097-4180. @article{weulersse_eomes-dependent_2020b,
title = {Eomes-Dependent Loss of the Co-activating Receptor CD226 Restrains CD8+ Ŧ Cell Anti-tumor Functions and Limits the Efficacy of Cancer Immunotherapy},
author = {Weulersse, Marianne and Asrir, Assia and Pichler, Andrea C. and Lemaitre, Lea and Braun, Matthias and Carrié, Nadège and Joubert, Marie-Véronique and Le Moine, Marie and Do Souto, Laura and Gaud, Guillaume and Das, Indrajit and Brauns, Elisa and Scarlata, Clara M. and Morandi, Elena and Sundarrajan, Ashmitha and Cuisinier, Marine and Buisson, Laure and Maheo, Sabrina and Kassem, Sahar and Agesta, Arantxa and Pérès, Michaël and Verhoeyen, Els and Martinez, Alejandra and Mazieres, Julien and Dupré, Loïc and Gossye, Thomas and Pancaldi, Vera and Guillerey, Camille and Ayyoub, Maha and Dejean, Anne S. and Saoudi, Abdelhadi and Goriely, Stanislas and Avet-Loiseau, Hervé and Bald, Tobias and Smyth, Mark J. and Martinet, Ludovic},
doi = {10.1016/j.immuni.2020.09.006},
issn = {1097-4180},
year = {2020},
date = {2020-01-01},
journal = {Immunity},
volume = {53},
number = {4},
pages = {824--839.e10},
abstract = {CD8+ T cells within the tumor microenvironment (TME) are exposed to various signals that ultimately determine functional outcomes. Here, we examined the role of the co-activating receptor CD226 (DNAM-1) in CD8+ T cell function. The absence of CD226 expression identified a subset of dysfunctional CD8+ T cells present in peripheral blood of healthy individuals. These cells exhibited reduced LFA-1 activation, altered TCR signaling, and a distinct transcriptomic program upon stimulation. CD226neg CD8+ T cells accumulated in human and mouse tumors of diverse origin through an antigen-specific mechanism involving the transcriptional regulator Eomesodermin (Eomes). Despite similar expression of co-inhibitory receptors, CD8+ tumor-infiltrating lymphocyte failed to respond to anti-PD-1 in the absence of CD226. Immune checkpoint blockade efficacy was hampered in Cd226-/- mice. Anti-CD137 (4-1BB) agonists also stimulated Eomes-dependent CD226 loss that limited the anti-tumor efficacy of this treatment. Thus, CD226 loss restrains CD8+ T cell function and limits the efficacy of cancer immunotherapy.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
CD8+ T cells within the tumor microenvironment (TME) are exposed to various signals that ultimately determine functional outcomes. Here, we examined the role of the co-activating receptor CD226 (DNAM-1) in CD8+ T cell function. The absence of CD226 expression identified a subset of dysfunctional CD8+ T cells present in peripheral blood of healthy individuals. These cells exhibited reduced LFA-1 activation, altered TCR signaling, and a distinct transcriptomic program upon stimulation. CD226neg CD8+ T cells accumulated in human and mouse tumors of diverse origin through an antigen-specific mechanism involving the transcriptional regulator Eomesodermin (Eomes). Despite similar expression of co-inhibitory receptors, CD8+ tumor-infiltrating lymphocyte failed to respond to anti-PD-1 in the absence of CD226. Immune checkpoint blockade efficacy was hampered in Cd226-/- mice. Anti-CD137 (4-1BB) agonists also stimulated Eomes-dependent CD226 loss that limited the anti-tumor efficacy of this treatment. Thus, CD226 loss restrains CD8+ T cell function and limits the efficacy of cancer immunotherapy. |
Blaize, Gaëtan; Daniels-Treffandier, Hélène; Aloulou, Meryem; Rouquié, Nelly; Yang, Cui; Marcellin, Marlène; Gador, Mylène; Benamar, Mehdi; Ducatez, Mariette; Song, Ki-Duk; Burlet-Schiltz, Odile; Saoudi, Abdelhadi; Love, Paul E.; Fazilleau, Nicolas; Gonzalez de Peredo, Anne; Lesourne, Renaud CD5 signalosome coordinates antagonist TCR signals to control the generation of Treg cells induced by foreign antigens Journal Article In: Proc Natl Acad Sci U S A, vol. 117, no. 23, pp. 12969–12979, 2020, ISSN: 1091-6490. @article{blaize_cd5_2020b,
title = {CD5 signalosome coordinates antagonist TCR signals to control the generation of Treg cells induced by foreign antigens},
author = {Blaize, Gaëtan and Daniels-Treffandier, Hélène and Aloulou, Meryem and Rouquié, Nelly and Yang, Cui and Marcellin, Marlène and Gador, Mylène and Benamar, Mehdi and Ducatez, Mariette and Song, Ki-Duk and Burlet-Schiltz, Odile and Saoudi, Abdelhadi and Love, Paul E. and Fazilleau, Nicolas and Gonzalez de Peredo, Anne and Lesourne, Renaud},
doi = {10.1073/pnas.1917182117},
issn = {1091-6490},
year = {2020},
date = {2020-01-01},
journal = {Proc Natl Acad Sci U S A},
volume = {117},
number = {23},
pages = {12969--12979},
abstract = {CD5 is characterized as an inhibitory coreceptor with an important regulatory role during T cell development. The molecular mechanism by which CD5 operates has been puzzling and its function in mature T cells suggests promoting rather than repressing effects on immune responses. Here, we combined quantitative mass spectrometry and genetic studies to analyze the components and the activity of the CD5 signaling machinery in primary T cells. We found that T cell receptor (TCR) engagement induces the selective phosphorylation of CD5 tyrosine 429, which serves as a docking site for proteins with adaptor functions (c-Cbl, CIN85, CRKL), connecting CD5 to positive (PI3K) and negative (UBASH3A, SHIP1) regulators of TCR signaling. c-CBL acts as a coordinator in this complex enabling CD5 to synchronize positive and negative feedbacks on TCR signaling through the other components. Disruption of CD5 signalosome in mutant mice reveals that it modulates TCR signal outputs to selectively repress the transactivation of Foxp3 and limit the inopportune induction of peripherally induced regulatory T cells during immune responses against foreign antigen. Our findings bring insights into the paradigm of coreceptor signaling, suggesting that, in addition to providing dualistic enhancing or dampening inputs, coreceptors can engage concomitant stimulatory and inhibitory signaling events, which act together to promote specific functional outcomes.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
CD5 is characterized as an inhibitory coreceptor with an important regulatory role during T cell development. The molecular mechanism by which CD5 operates has been puzzling and its function in mature T cells suggests promoting rather than repressing effects on immune responses. Here, we combined quantitative mass spectrometry and genetic studies to analyze the components and the activity of the CD5 signaling machinery in primary T cells. We found that T cell receptor (TCR) engagement induces the selective phosphorylation of CD5 tyrosine 429, which serves as a docking site for proteins with adaptor functions (c-Cbl, CIN85, CRKL), connecting CD5 to positive (PI3K) and negative (UBASH3A, SHIP1) regulators of TCR signaling. c-CBL acts as a coordinator in this complex enabling CD5 to synchronize positive and negative feedbacks on TCR signaling through the other components. Disruption of CD5 signalosome in mutant mice reveals that it modulates TCR signal outputs to selectively repress the transactivation of Foxp3 and limit the inopportune induction of peripherally induced regulatory T cells during immune responses against foreign antigen. Our findings bring insights into the paradigm of coreceptor signaling, suggesting that, in addition to providing dualistic enhancing or dampening inputs, coreceptors can engage concomitant stimulatory and inhibitory signaling events, which act together to promote specific functional outcomes. |
Yamak, Abir; Hu, Dongjian; Mittal, Nikhil; Buikema, Jan W.; Ditta, Sheraz; Lutz, Pierre G.; Moog-Lutz, Christel; Ellinor, Patrick T.; Domian, Ibrahim J. Loss of Asb2 Impairs Cardiomyocyte Differentiation and Leads to Congenital Double Outlet Right Ventricle Journal Article In: iScience, vol. 23, no. 3, pp. 100959, 2020, ISSN: 2589-0042. @article{yamak_loss_2020,
title = {Loss of Asb2 Impairs Cardiomyocyte Differentiation and Leads to Congenital Double Outlet Right Ventricle},
author = {Yamak, Abir and Hu, Dongjian and Mittal, Nikhil and Buikema, Jan W. and Ditta, Sheraz and Lutz, Pierre G. and Moog-Lutz, Christel and Ellinor, Patrick T. and Domian, Ibrahim J.},
doi = {10.1016/j.isci.2020.100959},
issn = {2589-0042},
year = {2020},
date = {2020-01-01},
journal = {iScience},
volume = {23},
number = {3},
pages = {100959},
abstract = {Defining the pathways that control cardiac development facilitates understanding the pathogenesis of congenital heart disease. Herein, we identify enrichment of a Cullin5 Ub ligase key subunit, Asb2, in myocardial progenitors and differentiated cardiomyocytes. Using two conditional murine knockouts, Nkx+/Cre.Asb2fl/fl and AHF-Cre.Asb2fl/fl, and tissue clarifying technique, we reveal Asb2 requirement for embryonic survival and complete heart looping. Deletion of Asb2 results in upregulation of its target Filamin A (Flna), and concurrent Flna deletion partially rescues embryonic lethality. Conditional AHF-Cre.Asb2 knockouts harboring one Flna allele have double outlet right ventricle (DORV), which is rescued by biallelic Flna excision. Transcriptomic and immunofluorescence analyses identify Tgfβ/Smad as downstream targets of Asb2/Flna. Finally, using CRISPR/Cas9 genome editing, we demonstrate Asb2 requirement for human cardiomyocyte differentiation suggesting a conserved mechanism between mice and humans. Collectively, our study provides deeper mechanistic understanding of the role of the ubiquitin proteasome system in cardiac development and suggests a previously unidentified murine model for DORV.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
Defining the pathways that control cardiac development facilitates understanding the pathogenesis of congenital heart disease. Herein, we identify enrichment of a Cullin5 Ub ligase key subunit, Asb2, in myocardial progenitors and differentiated cardiomyocytes. Using two conditional murine knockouts, Nkx+/Cre.Asb2fl/fl and AHF-Cre.Asb2fl/fl, and tissue clarifying technique, we reveal Asb2 requirement for embryonic survival and complete heart looping. Deletion of Asb2 results in upregulation of its target Filamin A (Flna), and concurrent Flna deletion partially rescues embryonic lethality. Conditional AHF-Cre.Asb2 knockouts harboring one Flna allele have double outlet right ventricle (DORV), which is rescued by biallelic Flna excision. Transcriptomic and immunofluorescence analyses identify Tgfβ/Smad as downstream targets of Asb2/Flna. Finally, using CRISPR/Cas9 genome editing, we demonstrate Asb2 requirement for human cardiomyocyte differentiation suggesting a conserved mechanism between mice and humans. Collectively, our study provides deeper mechanistic understanding of the role of the ubiquitin proteasome system in cardiac development and suggests a previously unidentified murine model for DORV. |
Salzer, Elisabeth; Zoghi, Samaneh; Kiss, Máté G.; Kage, Frieda; Rashkova, Christina; Stahnke, Stephanie; Haimel, Matthias; Platzer, René; Caldera, Michael; Ardy, Rico Chandra; Hoeger, Birgit; Block, Jana; Medgyesi, David; Sin, Celine; Shahkarami, Sepideh; Kain, Renate; Ziaee, Vahid; Hammerl, Peter; Bock, Christoph; Menche, Jörg; Dupré, Loïc; Huppa, Johannes B.; Sixt, Michael; Lomakin, Alexis; Rottner, Klemens; Binder, Christoph J.; Stradal, Theresia E. B.; Rezaei, Nima; Boztug, Kaan The cytoskeletal regulator HEM1 governs B cell development and prevents autoimmunity Journal Article In: Science Immunology, vol. 5, no. 49, 2020, ISSN: 2470-9468. @article{salzer_cytoskeletal_2020,
title = {The cytoskeletal regulator HEM1 governs B cell development and prevents autoimmunity},
author = {Salzer, Elisabeth and Zoghi, Samaneh and Kiss, Máté G. and Kage, Frieda and Rashkova, Christina and Stahnke, Stephanie and Haimel, Matthias and Platzer, René and Caldera, Michael and Ardy, Rico Chandra and Hoeger, Birgit and Block, Jana and Medgyesi, David and Sin, Celine and Shahkarami, Sepideh and Kain, Renate and Ziaee, Vahid and Hammerl, Peter and Bock, Christoph and Menche, Jörg and Dupré, Loïc and Huppa, Johannes B. and Sixt, Michael and Lomakin, Alexis and Rottner, Klemens and Binder, Christoph J. and Stradal, Theresia E. B. and Rezaei, Nima and Boztug, Kaan},
doi = {10.1126/sciimmunol.abc3979},
issn = {2470-9468},
year = {2020},
date = {2020-01-01},
journal = {Science Immunology},
volume = {5},
number = {49},
abstract = {The WAVE regulatory complex (WRC) is crucial for assembly of the peripheral branched actin network constituting one of the main drivers of eukaryotic cell migration. Here, we uncover an essential role of the hematopoietic-specific WRC component HEM1 for immune cell development. Germline-encoded HEM1 deficiency underlies an inborn error of immunity with systemic autoimmunity, at cellular level marked by WRC destabilization, reduced filamentous actin, and failure to assemble lamellipodia. Hem1-/- mice display systemic autoimmunity, phenocopying the human disease. In the absence of Hem1, B cells become deprived of extracellular stimuli necessary to maintain the strength of B cell receptor signaling at a level permissive for survival of non-autoreactive B cells. This shifts the balance of B cell fate choices toward autoreactive B cells and thus autoimmunity.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
The WAVE regulatory complex (WRC) is crucial for assembly of the peripheral branched actin network constituting one of the main drivers of eukaryotic cell migration. Here, we uncover an essential role of the hematopoietic-specific WRC component HEM1 for immune cell development. Germline-encoded HEM1 deficiency underlies an inborn error of immunity with systemic autoimmunity, at cellular level marked by WRC destabilization, reduced filamentous actin, and failure to assemble lamellipodia. Hem1-/- mice display systemic autoimmunity, phenocopying the human disease. In the absence of Hem1, B cells become deprived of extracellular stimuli necessary to maintain the strength of B cell receptor signaling at a level permissive for survival of non-autoreactive B cells. This shifts the balance of B cell fate choices toward autoreactive B cells and thus autoimmunity. |
Thian, Marini; Hoeger, Birgit; Kamnev, Anton; Poyer, Fiona; Köstel Bal, Sevgi; Caldera, Michael; Jiménez-Heredia, Raúl; Huemer, Jakob; Pickl, Winfried F.; Groß, Miriam; Ehl, Stephan; Lucas, Carrie L.; Menche, Jörg; Hutter, Caroline; Attarbaschi, Andishe; Dupré, Loïc; Boztug, Kaan Germline biallelic PIK3CG mutations in a multifaceted immunodeficiency with immune dysregulation Journal Article In: Haematologica, 2020, ISSN: 1592-8721. @article{thian_germline_2020b,
title = {Germline biallelic PIK3CG mutations in a multifaceted immunodeficiency with immune dysregulation},
author = {Thian, Marini and Hoeger, Birgit and Kamnev, Anton and Poyer, Fiona and Köstel Bal, Sevgi and Caldera, Michael and Jiménez-Heredia, Raúl and Huemer, Jakob and Pickl, Winfried F. and Groß, Miriam and Ehl, Stephan and Lucas, Carrie L. and Menche, Jörg and Hutter, Caroline and Attarbaschi, Andishe and Dupré, Loïc and Boztug, Kaan},
doi = {10.3324/haematol.2019.231399},
issn = {1592-8721},
year = {2020},
date = {2020-01-01},
journal = {Haematologica},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
|
Gallais, Fanny; Ysebaert, Loïc; Despas, Fabien; De Barros, Sandra; Dupré, Loïc; Quillet-Mary, Anne; Protin, Caroline; Thomas, Fabienne; Obéric, Lucie; Allal, Ben; Chatelut, Etienne; White-Koning, Mélanie Population Pharmacokinetics of Ibrutinib and Its Dihydrodiol Metabolite in Patients with Lymphoid Malignancies Journal Article In: Clinical Pharmacokinetics, 2020, ISSN: 1179-1926. @article{gallais_population_2020b,
title = {Population Pharmacokinetics of Ibrutinib and Its Dihydrodiol Metabolite in Patients with Lymphoid Malignancies},
author = {Gallais, Fanny and Ysebaert, Loïc and Despas, Fabien and De Barros, Sandra and Dupré, Loïc and Quillet-Mary, Anne and Protin, Caroline and Thomas, Fabienne and Obéric, Lucie and Allal, Ben and Chatelut, Etienne and White-Koning, Mélanie},
doi = {10.1007/s40262-020-00884-0},
issn = {1179-1926},
year = {2020},
date = {2020-01-01},
journal = {Clinical Pharmacokinetics},
abstract = {BACKGROUND AND OBJECTIVE: Ibrutinib is used for the treatment of chronic lymphocytic leukemia and other lymphoid malignancies. The aim of this work is to develop a population pharmacokinetic model for ibrutinib and its dihydrodiol metabolite to quantify pharmacokinetic inter- and intra-individual variability, to evaluate the impact of several covariates on ibrutinib pharmacokinetic parameters, and to examine the relationship between exposure and clinical outcome.
METHODS: Patients treated with ibrutinib were included in the study and followed up for 2 years. Pharmacokinetic blood samples were taken from months 1 to 12 after inclusion. Ibrutinib and dihydrodiol-ibrutinib concentrations were assessed using ultra-performance liquid chromatography tandem mass spectrometry. A population pharmacokinetic model was developed using NONMEM version 7.4. RESULTS: A total of 89 patients and 1501 plasma concentrations were included in the pharmacokinetic analysis. The best model consisted in two compartments for each molecule. Absorption was described by a sequential zero first-order process and a lag time. Ibrutinib was either metabolised into dihydrodiol-ibrutinib or excreted through other elimination routes. A link between the dosing compartment and the dihydrodiol-ibrutinib central compartment was added to assess for high first-pass hepatic metabolism. Ibrutinib clearance had 67% and 47% inter- and intra-individual variability, respectively, while dihydrodiol-ibrutinib clearance had 51% and 26% inter- and intra-individual variability, respectively. Observed ibrutinib exposure is significantly higher in patients carrying one copy of the cytochrome P450 3A4*22 variant (1167 ng.h/mL vs 743 ng.h/mL, respectively},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
BACKGROUND AND OBJECTIVE: Ibrutinib is used for the treatment of chronic lymphocytic leukemia and other lymphoid malignancies. The aim of this work is to develop a population pharmacokinetic model for ibrutinib and its dihydrodiol metabolite to quantify pharmacokinetic inter- and intra-individual variability, to evaluate the impact of several covariates on ibrutinib pharmacokinetic parameters, and to examine the relationship between exposure and clinical outcome.
METHODS: Patients treated with ibrutinib were included in the study and followed up for 2 years. Pharmacokinetic blood samples were taken from months 1 to 12 after inclusion. Ibrutinib and dihydrodiol-ibrutinib concentrations were assessed using ultra-performance liquid chromatography tandem mass spectrometry. A population pharmacokinetic model was developed using NONMEM version 7.4. RESULTS: A total of 89 patients and 1501 plasma concentrations were included in the pharmacokinetic analysis. The best model consisted in two compartments for each molecule. Absorption was described by a sequential zero first-order process and a lag time. Ibrutinib was either metabolised into dihydrodiol-ibrutinib or excreted through other elimination routes. A link between the dosing compartment and the dihydrodiol-ibrutinib central compartment was added to assess for high first-pass hepatic metabolism. Ibrutinib clearance had 67% and 47% inter- and intra-individual variability, respectively, while dihydrodiol-ibrutinib clearance had 51% and 26% inter- and intra-individual variability, respectively. Observed ibrutinib exposure is significantly higher in patients carrying one copy of the cytochrome P450 3A4*22 variant (1167 ng.h/mL vs 743 ng.h/mL, respectively |
2019
|
Spinner, Camille A.; Lamsoul, Isabelle; Métais, Arnaud; Febrissy, Chanaëlle; Moog-Lutz, Christel; Lutz, Pierre G. The E3 Ubiquitin Ligase Asb2α in Ŧ Helper 2 Cells Negatively Regulates Antitumor Immunity in Colorectal Cancer Journal Article In: Cancer Immunol Res, vol. 7, no. 8, pp. 1332–1344, 2019, ISSN: 2326-6074. @article{spinner_e3_2019,
title = {The E3 Ubiquitin Ligase Asb2α in Ŧ Helper 2 Cells Negatively Regulates Antitumor Immunity in Colorectal Cancer},
author = {Spinner, Camille A. and Lamsoul, Isabelle and Métais, Arnaud and Febrissy, Chanaëlle and Moog-Lutz, Christel and Lutz, Pierre G.},
doi = {10.1158/2326-6066.CIR-18-0562},
issn = {2326-6074},
year = {2019},
date = {2019-01-01},
journal = {Cancer Immunol Res},
volume = {7},
number = {8},
pages = {1332--1344},
abstract = {The escape of cancer cells from host immunosurveillance involves a shift in immune responses, including an imbalance in Th1 and Th2 cells. A Th1-dominated immune response predicts positive outcomes in colorectal cancer. The E3 ubiquitin ligase, Asb2α, is expressed in Th2 cells, but its roles in T-cell maturation and cancer are unclear. We provide evidence that the Th2 master regulator, Gata3, induces Asb2 Loss of Asb2 did not affect Th differentiation ex vivo, but reduced IL4 production from Th2 cells. We found that high ASB2 expression was associated with poor outcome in colorectal cancer. Loss of Asb2 from hematopoietic cells promoted a Th1 response and attenuated colitis-associated tumorigenesis in mice. Diminished Th2 function correlated with increased IFNγ production and an enhanced type 1 antitumor immune response in Asb2-deficient mice. Our work suggests that Asb2α promotes a Th2 phenotype in vivo, which in turn is associated with tumor progression in a mouse model of colitis.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
The escape of cancer cells from host immunosurveillance involves a shift in immune responses, including an imbalance in Th1 and Th2 cells. A Th1-dominated immune response predicts positive outcomes in colorectal cancer. The E3 ubiquitin ligase, Asb2α, is expressed in Th2 cells, but its roles in T-cell maturation and cancer are unclear. We provide evidence that the Th2 master regulator, Gata3, induces Asb2 Loss of Asb2 did not affect Th differentiation ex vivo, but reduced IL4 production from Th2 cells. We found that high ASB2 expression was associated with poor outcome in colorectal cancer. Loss of Asb2 from hematopoietic cells promoted a Th1 response and attenuated colitis-associated tumorigenesis in mice. Diminished Th2 function correlated with increased IFNγ production and an enhanced type 1 antitumor immune response in Asb2-deficient mice. Our work suggests that Asb2α promotes a Th2 phenotype in vivo, which in turn is associated with tumor progression in a mouse model of colitis. |
Cheminant, Morgane; Mahlaoui, Nizar; Desconclois, Céline; Canioni, Danielle; Ysebaert, Loïc; Dupré, Loïc; Vasconcelos, Zilton; Malphettes, Marion; Moshous, Despina; Neven, Bénédicte; Rohrlich, Pierre-Simon; Bernard, Marc; Bertrand, Yves; Fischer, Alain; Suarez, Felipe Lymphoproliferative disease in patients with Wiskott-Aldrich syndrome: Analysis of the French Registry of Primary Immunodeficiencies Journal Article In: The Journal of Allergy and Clinical Immunology, vol. 143, no. 6, pp. 2311–2315.e7, 2019, ISSN: 1097-6825. @article{cheminant_lymphoproliferative_2019b,
title = {Lymphoproliferative disease in patients with Wiskott-Aldrich syndrome: Analysis of the French Registry of Primary Immunodeficiencies},
author = {Cheminant, Morgane and Mahlaoui, Nizar and Desconclois, Céline and Canioni, Danielle and Ysebaert, Loïc and Dupré, Loïc and Vasconcelos, Zilton and Malphettes, Marion and Moshous, Despina and Neven, Bénédicte and Rohrlich, Pierre-Simon and Bernard, Marc and Bertrand, Yves and Fischer, Alain and Suarez, Felipe},
doi = {10.1016/j.jaci.2019.01.046},
issn = {1097-6825},
year = {2019},
date = {2019-01-01},
journal = {The Journal of Allergy and Clinical Immunology},
volume = {143},
number = {6},
pages = {2311--2315.e7},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
|
Guipouy, Delphine; Gertner-Dardenne, Julie; Pfajfer, Laurène; German, Yolla; Belmonte, Nathalie; Dupré, Loïc Granulysin- and granzyme-dependent elimination of myeloid cells by therapeutic ova-specific type 1 regulatory Ŧ cells Journal Article In: International Immunology, vol. 31, no. 4, pp. 239–250, 2019, ISSN: 1460-2377. @article{guipouy_granulysin-_2019,
title = {Granulysin- and granzyme-dependent elimination of myeloid cells by therapeutic ova-specific type 1 regulatory Ŧ cells},
author = {Guipouy, Delphine and Gertner-Dardenne, Julie and Pfajfer, Laurène and German, Yolla and Belmonte, Nathalie and Dupré, Loïc},
doi = {10.1093/intimm/dxy083},
issn = {1460-2377},
year = {2019},
date = {2019-01-01},
journal = {International Immunology},
volume = {31},
number = {4},
pages = {239--250},
abstract = {The intrinsic immunosuppressive properties of regulatory T (Treg) cells can be harnessed for therapeutic approaches aiming at down-modulating harmful immune reactions. In this context, expanded type 1 Treg cells (Tr1 cells) specific for ovalbumin (ova-Tr1 cells) have been tested for clinical efficacy in the treatment of autoimmune disorders such as refractory Crohn's disease (CD). The clinical use of these therapeutic products warrants exploration of their mechanism of action. Here, we identified a relationship between the CD activity index and the expression of lytic molecules by the ova-Tr1 cells administered in the previously reported First-in-Man study [Crohn's And Treg cells Study 1 (CATS1) study]. Accordingly, ova-Tr1 cells were found to carry granules containing high levels of lytic molecules, including multiple granzymes and granulysin. These cells displayed a T-cell receptor (TCR)-independent cytotoxic activity, which was preferentially directed toward myeloid cell lines and monocyte-derived dendritic cells. Upon contact with myeloid cells, ova-Tr1 cells induced their apoptosis via a perforin-independent and a granulysin/granzyme-dependent mechanism. As compared to CD8+ cytotoxic T cells, ova-Tr1 cells required more time to lyse target cells and displayed a more gradual lytic activity over time. Notably, this activity was sustained over days resulting in the control of myeloid cell populations at a relatively low ratio. Our study reveals that ova-Tr1 cells are endowed with a sustained cytotoxic activity that relies on a unique combination of granulysin and granzymes and that preferentially eliminates myeloid target cells in a TCR-independent manner.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
The intrinsic immunosuppressive properties of regulatory T (Treg) cells can be harnessed for therapeutic approaches aiming at down-modulating harmful immune reactions. In this context, expanded type 1 Treg cells (Tr1 cells) specific for ovalbumin (ova-Tr1 cells) have been tested for clinical efficacy in the treatment of autoimmune disorders such as refractory Crohn's disease (CD). The clinical use of these therapeutic products warrants exploration of their mechanism of action. Here, we identified a relationship between the CD activity index and the expression of lytic molecules by the ova-Tr1 cells administered in the previously reported First-in-Man study [Crohn's And Treg cells Study 1 (CATS1) study]. Accordingly, ova-Tr1 cells were found to carry granules containing high levels of lytic molecules, including multiple granzymes and granulysin. These cells displayed a T-cell receptor (TCR)-independent cytotoxic activity, which was preferentially directed toward myeloid cell lines and monocyte-derived dendritic cells. Upon contact with myeloid cells, ova-Tr1 cells induced their apoptosis via a perforin-independent and a granulysin/granzyme-dependent mechanism. As compared to CD8+ cytotoxic T cells, ova-Tr1 cells required more time to lyse target cells and displayed a more gradual lytic activity over time. Notably, this activity was sustained over days resulting in the control of myeloid cell populations at a relatively low ratio. Our study reveals that ova-Tr1 cells are endowed with a sustained cytotoxic activity that relies on a unique combination of granulysin and granzymes and that preferentially eliminates myeloid target cells in a TCR-independent manner. |
2018
|
Gaud, G.; Lesourne, R.; Love, P. E. Regulatory mechanisms in T cell receptor signalling Journal Article In: Nat Rev Immunol, 2018, ISSN: 1474-1741 (Electronic)
1474-1733 (Linking). @article{RN1b,
title = {Regulatory mechanisms in T cell receptor signalling},
author = {Gaud, G. and Lesourne, R. and Love, P. E.},
url = {https://www.ncbi.nlm.nih.gov/pubmed/29789755},
doi = {10.1038/s41577-018-0020-8},
issn = {1474-1741 (Electronic)
1474-1733 (Linking)},
year = {2018},
date = {2018-01-01},
journal = {Nat Rev Immunol},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
|
Houmadi, R.; Guipouy, D.; Rey-Barroso, J.; Vasconcelos, Z.; Cornet, J.; Manghi, M.; Destainville, N.; Valitutti, S.; Allart, S.; Dupre, L. The Wiskott-Aldrich Syndrome Protein Contributes to the Assembly of the LFA-1 Nanocluster Belt at the Lytic Synapse Journal Article In: Cell Rep, vol. 22, no. 4, pp. 979-991, 2018, ISSN: 2211-1247 (Electronic). @article{RN3b,
title = {The Wiskott-Aldrich Syndrome Protein Contributes to the Assembly of the LFA-1 Nanocluster Belt at the Lytic Synapse},
author = {Houmadi, R. and Guipouy, D. and Rey-Barroso, J. and Vasconcelos, Z. and Cornet, J. and Manghi, M. and Destainville, N. and Valitutti, S. and Allart, S. and Dupre, L.},
url = {https://www.ncbi.nlm.nih.gov/pubmed/29386139},
doi = {10.1016/j.celrep.2017.12.088},
issn = {2211-1247 (Electronic)},
year = {2018},
date = {2018-01-01},
journal = {Cell Rep},
volume = {22},
number = {4},
pages = {979-991},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
|
Rey-Barroso, Javier; Calovi, Daniel S.; Combe, Maud; German, Yolla; Moreau, Mathieu; Canivet, Astrid; Wang, Xiaobo; Sire, Clément; Theraulaz, Guy; Dupré, Loïc Switching between individual and collective motility in B lymphocytes is controlled by cell-matrix adhesion and inter-cellular interactions Journal Article In: Scientific Reports, vol. 8, no. 1, pp. 5800, 2018, ISSN: 2045-2322. @article{rey-barroso_switching_2018,
title = {Switching between individual and collective motility in B lymphocytes is controlled by cell-matrix adhesion and inter-cellular interactions},
author = { Javier Rey-Barroso and Daniel S. Calovi and Maud Combe and Yolla German and Mathieu Moreau and Astrid Canivet and Xiaobo Wang and Clément Sire and Guy Theraulaz and Loïc Dupré},
doi = {10.1038/s41598-018-24222-4},
issn = {2045-2322},
year = {2018},
date = {2018-01-01},
journal = {Scientific Reports},
volume = {8},
number = {1},
pages = {5800},
abstract = {Lymphocytes alternate between phases of individual migration across tissues and phases of clustering during activation and function. The range of lymphocyte motility behaviors and the identity of the factors that govern them remain elusive. To explore this point, we here collected unprecedented statistics pertaining to cell displacements, cell:matrix and cell:cell interactions using a model B cell line as well as primary human B lymphocytes. At low cell density, individual B lymphocytes displayed a high heterogeneity in their speed and diffusivity. Beyond this intrinsic variability, B lymphocytes adapted their motility to the composition of extra-cellular matrix, adopting slow persistent walks over collagen IV and quick Brownian walks over fibronectin. At high cell density, collagen IV favored the self-assembly of B lymphocytes into clusters endowed with collective coordination, while fibronectin stimulated individual motility. We show that this behavioral plasticity is controlled by acto-myosin dependent adhesive and Arp2/3-dependent protrusive actin pools, respectively. Our study reveals the adaptive nature of B lymphocyte motility and group dynamics, which are shaped by an interplay between and cell:matrix and cell:cell interactions.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
Lymphocytes alternate between phases of individual migration across tissues and phases of clustering during activation and function. The range of lymphocyte motility behaviors and the identity of the factors that govern them remain elusive. To explore this point, we here collected unprecedented statistics pertaining to cell displacements, cell:matrix and cell:cell interactions using a model B cell line as well as primary human B lymphocytes. At low cell density, individual B lymphocytes displayed a high heterogeneity in their speed and diffusivity. Beyond this intrinsic variability, B lymphocytes adapted their motility to the composition of extra-cellular matrix, adopting slow persistent walks over collagen IV and quick Brownian walks over fibronectin. At high cell density, collagen IV favored the self-assembly of B lymphocytes into clusters endowed with collective coordination, while fibronectin stimulated individual motility. We show that this behavioral plasticity is controlled by acto-myosin dependent adhesive and Arp2/3-dependent protrusive actin pools, respectively. Our study reveals the adaptive nature of B lymphocyte motility and group dynamics, which are shaped by an interplay between and cell:matrix and cell:cell interactions. |
Sarkar, Koustav; Han, Seong-Su; Wen, Kuo-Kuang; Ochs, Hans D.; Dupré, Loïc; Seidman, Michael M.; Vyas, Yatin M. R-loops cause genomic instability in Ŧ helper lymphocytes from patients with Wiskott-Aldrich syndrome Journal Article In: The Journal of Allergy and Clinical Immunology, vol. 142, no. 1, pp. 219–234, 2018, ISSN: 1097-6825. @article{sarkar_r-loops_2018,
title = {R-loops cause genomic instability in Ŧ helper lymphocytes from patients with Wiskott-Aldrich syndrome},
author = {Sarkar, Koustav and Han, Seong-Su and Wen, Kuo-Kuang and Ochs, Hans D. and Dupré, Loïc and Seidman, Michael M. and Vyas, Yatin M.},
doi = {10.1016/j.jaci.2017.11.023},
issn = {1097-6825},
year = {2018},
date = {2018-01-01},
journal = {The Journal of Allergy and Clinical Immunology},
volume = {142},
number = {1},
pages = {219--234},
abstract = {BACKGROUND: Wiskott-Aldrich syndrome (WAS), X-linked thrombocytopenia (XLT), and X-linked neutropenia, which are caused by WAS mutations affecting Wiskott-Aldrich syndrome protein (WASp) expression or activity, manifest in immunodeficiency, autoimmunity, genomic instability, and lymphoid and other cancers. WASp supports filamentous actin formation in the cytoplasm and gene transcription in the nucleus. Although the genetic basis for XLT/WAS has been clarified, the relationships between mutant forms of WASp and the diverse features of these disorders remain ill-defined.
OBJECTIVE: We sought to define how dysfunctional gene transcription is causally linked to the degree of TH cell deficiency and genomic instability in the XLT/WAS clinical spectrum.
METHODS: In human TH1- or TH2-skewing cell culture systems, cotranscriptional R-loops (RNA/DNA duplex and displaced single-stranded DNA) and DNA double-strand breaks (DSBs) were monitored in multiple samples from patients with XLT and WAS and in normal T cells depleted of WASp.
RESULTS: WASp deficiency provokes increased R-loops and R-loop-mediated DSBs in TH1 cells relative to TH2 cells. Mechanistically, chromatin occupancy of serine 2-unphosphorylated RNA polymerase II is increased, and that of topoisomerase 1, an R-loop preventing factor, is decreased at R-loop-enriched regions of IFNG and TBX21 (TH1 genes) in TH1 cells. These aberrations accompany increased unspliced (intron-retained) and decreased spliced mRNA of IFNG and TBX21 but not IL13 (TH2 gene). Significantly, increased cellular load of R-loops and DSBs, which are normalized on RNaseH1-mediated suppression of ectopic R-loops, inversely correlates with disease severity scores.
CONCLUSION: Transcriptional R-loop imbalance is a novel molecular defect causative in TH1 immunodeficiency and genomic instability in patients with WAS. The study proposes that cellular R-loop load could be used as a potential biomarker for monitoring symptom severity and prognostic outcome in the XLT-WAS clinical spectrum and could be targeted therapeutically.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
BACKGROUND: Wiskott-Aldrich syndrome (WAS), X-linked thrombocytopenia (XLT), and X-linked neutropenia, which are caused by WAS mutations affecting Wiskott-Aldrich syndrome protein (WASp) expression or activity, manifest in immunodeficiency, autoimmunity, genomic instability, and lymphoid and other cancers. WASp supports filamentous actin formation in the cytoplasm and gene transcription in the nucleus. Although the genetic basis for XLT/WAS has been clarified, the relationships between mutant forms of WASp and the diverse features of these disorders remain ill-defined.
OBJECTIVE: We sought to define how dysfunctional gene transcription is causally linked to the degree of TH cell deficiency and genomic instability in the XLT/WAS clinical spectrum.
METHODS: In human TH1- or TH2-skewing cell culture systems, cotranscriptional R-loops (RNA/DNA duplex and displaced single-stranded DNA) and DNA double-strand breaks (DSBs) were monitored in multiple samples from patients with XLT and WAS and in normal T cells depleted of WASp.
RESULTS: WASp deficiency provokes increased R-loops and R-loop-mediated DSBs in TH1 cells relative to TH2 cells. Mechanistically, chromatin occupancy of serine 2-unphosphorylated RNA polymerase II is increased, and that of topoisomerase 1, an R-loop preventing factor, is decreased at R-loop-enriched regions of IFNG and TBX21 (TH1 genes) in TH1 cells. These aberrations accompany increased unspliced (intron-retained) and decreased spliced mRNA of IFNG and TBX21 but not IL13 (TH2 gene). Significantly, increased cellular load of R-loops and DSBs, which are normalized on RNaseH1-mediated suppression of ectopic R-loops, inversely correlates with disease severity scores.
CONCLUSION: Transcriptional R-loop imbalance is a novel molecular defect causative in TH1 immunodeficiency and genomic instability in patients with WAS. The study proposes that cellular R-loop load could be used as a potential biomarker for monitoring symptom severity and prognostic outcome in the XLT-WAS clinical spectrum and could be targeted therapeutically. |
Pfajfer, Laurène; Mair, Nina K.; Jiménez-Heredia, Raúl; Genel, Ferah; Gulez, Nesrin; Ardeniz, Ömür; Hoeger, Birgit; Bal, Sevgi Köstel; Madritsch, Christoph; Kalinichenko, Artem; Chandra Ardy, Rico; Gerçeker, Bengü; Rey-Barroso, Javier; Ijspeert, Hanna; Tangye, Stuart G.; Simonitsch-Klupp, Ingrid; Huppa, Johannes B.; van der Burg, Mirjam; Dupré, Loïc; Boztug, Kaan Mutations affecting the actin regulator WD repeat-containing protein 1 lead to aberrant lymphoid immunity Journal Article In: The Journal of Allergy and Clinical Immunology, vol. 142, no. 5, pp. 1589–1604.e11, 2018, ISSN: 1097-6825. @article{pfajfer_mutations_2018,
title = {Mutations affecting the actin regulator WD repeat-containing protein 1 lead to aberrant lymphoid immunity},
author = {Pfajfer, Laurène and Mair, Nina K. and Jiménez-Heredia, Raúl and Genel, Ferah and Gulez, Nesrin and Ardeniz, Ömür and Hoeger, Birgit and Bal, Sevgi Köstel and Madritsch, Christoph and Kalinichenko, Artem and Chandra Ardy, Rico and Gerçeker, Bengü and Rey-Barroso, Javier and Ijspeert, Hanna and Tangye, Stuart G. and Simonitsch-Klupp, Ingrid and Huppa, Johannes B. and van der Burg, Mirjam and Dupré, Loïc and Boztug, Kaan},
doi = {10.1016/j.jaci.2018.04.023},
issn = {1097-6825},
year = {2018},
date = {2018-01-01},
journal = {The Journal of Allergy and Clinical Immunology},
volume = {142},
number = {5},
pages = {1589--1604.e11},
abstract = {BACKGROUND: The actin-interacting protein WD repeat-containing protein 1 (WDR1) promotes cofilin-dependent actin filament turnover. Biallelic WDR1 mutations have been identified recently in an immunodeficiency/autoinflammatory syndrome with aberrant morphology and function of myeloid cells.
OBJECTIVE: Given the pleiotropic expression of WDR1, here we investigated to what extent it might control the lymphoid arm of the immune system in human subjects.
METHODS: Histologic and detailed immunologic analyses were performed to elucidate the role of WDR1 in the development and function of B and T lymphocytes.
RESULTS: Here we identified novel homozygous and compound heterozygous WDR1 missense mutations in 6 patients belonging to 3 kindreds who presented with respiratory tract infections, skin ulceration, and stomatitis. In addition to defective adhesion and motility of neutrophils and monocytes, WDR1 deficiency was associated with aberrant T-cell activation and B-cell development. T lymphocytes appeared to develop normally in the patients, except for the follicular helper T-cell subset. However, peripheral T cells from the patients accumulated atypical actin structures at the immunologic synapse and displayed reduced calcium flux and mildly impaired proliferation on T-cell receptor stimulation. WDR1 deficiency was associated with even more severe abnormalities of the B-cell compartment, including peripheral B-cell lymphopenia, paucity of B-cell progenitors in the bone marrow, lack of switched memory B cells, reduced clonal diversity, abnormal B-cell spreading, and increased apoptosis on B-cell receptor/Toll-like receptor stimulation.
CONCLUSION: Our study identifies a novel role for WDR1 in adaptive immunity, highlighting WDR1 as a central regulator of actin turnover during formation of the B-cell and T-cell immunologic synapses.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
BACKGROUND: The actin-interacting protein WD repeat-containing protein 1 (WDR1) promotes cofilin-dependent actin filament turnover. Biallelic WDR1 mutations have been identified recently in an immunodeficiency/autoinflammatory syndrome with aberrant morphology and function of myeloid cells.
OBJECTIVE: Given the pleiotropic expression of WDR1, here we investigated to what extent it might control the lymphoid arm of the immune system in human subjects.
METHODS: Histologic and detailed immunologic analyses were performed to elucidate the role of WDR1 in the development and function of B and T lymphocytes.
RESULTS: Here we identified novel homozygous and compound heterozygous WDR1 missense mutations in 6 patients belonging to 3 kindreds who presented with respiratory tract infections, skin ulceration, and stomatitis. In addition to defective adhesion and motility of neutrophils and monocytes, WDR1 deficiency was associated with aberrant T-cell activation and B-cell development. T lymphocytes appeared to develop normally in the patients, except for the follicular helper T-cell subset. However, peripheral T cells from the patients accumulated atypical actin structures at the immunologic synapse and displayed reduced calcium flux and mildly impaired proliferation on T-cell receptor stimulation. WDR1 deficiency was associated with even more severe abnormalities of the B-cell compartment, including peripheral B-cell lymphopenia, paucity of B-cell progenitors in the bone marrow, lack of switched memory B cells, reduced clonal diversity, abnormal B-cell spreading, and increased apoptosis on B-cell receptor/Toll-like receptor stimulation.
CONCLUSION: Our study identifies a novel role for WDR1 in adaptive immunity, highlighting WDR1 as a central regulator of actin turnover during formation of the B-cell and T-cell immunologic synapses. |
Métais, Arnaud; Lamsoul, Isabelle; Melet, Armelle; Uttenweiler-Joseph, Sandrine; Poincloux, Renaud; Stefanovic, Sonia; Valière, Amélie; Gonzalez de Peredo, Anne; Stella, Alexandre; Burlet-Schiltz, Odile; Zaffran, Stéphane; Lutz, Pierre G.; Moog-Lutz, Christel Asb2α-Filamin A Axis Is Essential for Actin Cytoskeleton Remodeling During Heart Development Journal Article In: Circ. Res., vol. 122, no. 6, pp. e34–e48, 2018, ISSN: 1524-4571. @article{metais_asb2-filamin_2018,
title = {Asb2α-Filamin A Axis Is Essential for Actin Cytoskeleton Remodeling During Heart Development},
author = {Métais, Arnaud and Lamsoul, Isabelle and Melet, Armelle and Uttenweiler-Joseph, Sandrine and Poincloux, Renaud and Stefanovic, Sonia and Valière, Amélie and Gonzalez de Peredo, Anne and Stella, Alexandre and Burlet-Schiltz, Odile and Zaffran, Stéphane and Lutz, Pierre G. and Moog-Lutz, Christel},
doi = {10.1161/CIRCRESAHA.117.312015},
issn = {1524-4571},
year = {2018},
date = {2018-01-01},
journal = {Circ. Res.},
volume = {122},
number = {6},
pages = {e34--e48},
abstract = {RATIONALE: Heart development involves differentiation of cardiac progenitors and assembly of the contractile sarcomere apparatus of cardiomyocytes. However, little is known about the mechanisms that regulate actin cytoskeleton remodeling during cardiac cell differentiation.
OBJECTIVE: The Asb2α (Ankyrin repeat-containing protein with a suppressor of cytokine signaling box 2) CRL5 (cullin 5 RING E3 ubiquitin ligase) triggers polyubiquitylation and subsequent degradation by the proteasome of FLNs (filamins). Here, we investigate the role of Asb2α in heart development and its mechanisms of action.
METHODS AND RESULTS: Using Asb2 knockout embryos, we show that Asb2 is an essential gene, critical to heart morphogenesis and function, although its loss does not interfere with the overall patterning of the embryonic heart tube. We show that the Asb2α E3 ubiquitin ligase controls Flna stability in immature cardiomyocytes. Importantly, Asb2α-mediated degradation of the actin-binding protein Flna marks a previously unrecognized intermediate step in cardiac cell differentiation characterized by cell shape changes and actin cytoskeleton remodeling. We further establish that in the absence of Asb2α, myofibrils are disorganized and that heartbeats are inefficient, leading to embryonic lethality in mice.
CONCLUSIONS: These findings identify Asb2α as an unsuspected key regulator of cardiac cell differentiation and shed light on the molecular and cellular mechanisms determining the onset of myocardial cell architecture and its link with early cardiac function. Although Flna is known to play roles in cytoskeleton organization and to be required for heart function, this study now reveals that its degradation mediated by Asb2α ensures essential functions in differentiating cardiac progenitors.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
RATIONALE: Heart development involves differentiation of cardiac progenitors and assembly of the contractile sarcomere apparatus of cardiomyocytes. However, little is known about the mechanisms that regulate actin cytoskeleton remodeling during cardiac cell differentiation.
OBJECTIVE: The Asb2α (Ankyrin repeat-containing protein with a suppressor of cytokine signaling box 2) CRL5 (cullin 5 RING E3 ubiquitin ligase) triggers polyubiquitylation and subsequent degradation by the proteasome of FLNs (filamins). Here, we investigate the role of Asb2α in heart development and its mechanisms of action.
METHODS AND RESULTS: Using Asb2 knockout embryos, we show that Asb2 is an essential gene, critical to heart morphogenesis and function, although its loss does not interfere with the overall patterning of the embryonic heart tube. We show that the Asb2α E3 ubiquitin ligase controls Flna stability in immature cardiomyocytes. Importantly, Asb2α-mediated degradation of the actin-binding protein Flna marks a previously unrecognized intermediate step in cardiac cell differentiation characterized by cell shape changes and actin cytoskeleton remodeling. We further establish that in the absence of Asb2α, myofibrils are disorganized and that heartbeats are inefficient, leading to embryonic lethality in mice.
CONCLUSIONS: These findings identify Asb2α as an unsuspected key regulator of cardiac cell differentiation and shed light on the molecular and cellular mechanisms determining the onset of myocardial cell architecture and its link with early cardiac function. Although Flna is known to play roles in cytoskeleton organization and to be required for heart function, this study now reveals that its degradation mediated by Asb2α ensures essential functions in differentiating cardiac progenitors. |
Brigida, Immacolata; Zoccolillo, Matteo; Cicalese, Maria Pia; Pfajfer, Laurène; Barzaghi, Federica; Scala, Serena; Oleaga-Quintas, Carmen; Álvarez-Álvarez, Jesus A.; Sereni, Lucia; Giannelli, Stefania; Sartirana, Claudia; Dionisio, Francesca; Pavesi, Luca; Benavides-Nieto, Marta; Basso-Ricci, Luca; Capasso, Paola; Mazzi, Benedetta; Rosain, Jeremie; Marcus, Nufar; Lee, Yu Nee; Somech, Raz; Degano, Massimo; Raiola, Giuseppe; Caorsi, Roberta; Picco, Paolo; Moncada Velez, Marcela; Khourieh, Joelle; Arias, Andrés Augusto; Bousfiha, Aziz; Issekutz, Thomas; Issekutz, Andrew; Boisson, Bertrand; Dobbs, Kerry; Villa, Anna; Lombardo, Angelo; Neven, Benedicte; Moshous, Despina; Casanova, Jean-Laurent; Franco, José Luis; Notarangelo, Luigi D.; Scielzo, Cristina; Volpi, Stefano; Dupré, Loïc; Bustamante, Jacinta; Gattorno, Marco; Aiuti, Alessandro T-cell defects in patients with ARPC1B germline mutations account for combined immunodeficiency Journal Article In: Blood, vol. 132, no. 22, pp. 2362–2374, 2018, ISSN: 1528-0020. @article{brigida_t-cell_2018,
title = {T-cell defects in patients with ARPC1B germline mutations account for combined immunodeficiency},
author = {Brigida, Immacolata and Zoccolillo, Matteo and Cicalese, Maria Pia and Pfajfer, Laurène and Barzaghi, Federica and Scala, Serena and Oleaga-Quintas, Carmen and Álvarez-Álvarez, Jesus A. and Sereni, Lucia and Giannelli, Stefania and Sartirana, Claudia and Dionisio, Francesca and Pavesi, Luca and Benavides-Nieto, Marta and Basso-Ricci, Luca and Capasso, Paola and Mazzi, Benedetta and Rosain, Jeremie and Marcus, Nufar and Lee, Yu Nee and Somech, Raz and Degano, Massimo and Raiola, Giuseppe and Caorsi, Roberta and Picco, Paolo and Moncada Velez, Marcela and Khourieh, Joelle and Arias, Andrés Augusto and Bousfiha, Aziz and Issekutz, Thomas and Issekutz, Andrew and Boisson, Bertrand and Dobbs, Kerry and Villa, Anna and Lombardo, Angelo and Neven, Benedicte and Moshous, Despina and Casanova, Jean-Laurent and Franco, José Luis and Notarangelo, Luigi D. and Scielzo, Cristina and Volpi, Stefano and Dupré, Loïc and Bustamante, Jacinta and Gattorno, Marco and Aiuti, Alessandro},
doi = {10.1182/blood-2018-07-863431},
issn = {1528-0020},
year = {2018},
date = {2018-01-01},
journal = {Blood},
volume = {132},
number = {22},
pages = {2362--2374},
abstract = {ARPC1B is a key factor for the assembly and maintenance of the ARP2/3 complex that is involved in actin branching from an existing filament. Germline biallelic mutations in ARPC1B have been recently described in 6 patients with clinical features of combined immunodeficiency (CID), whose neutrophils and platelets but not T lymphocytes were studied. We hypothesized that ARPC1B deficiency may also lead to cytoskeleton and functional defects in T cells. We have identified biallelic mutations in ARPC1B in 6 unrelated patients with early onset disease characterized by severe infections, autoimmune manifestations, and thrombocytopenia. Immunological features included T-cell lymphopenia, low numbers of naïve T cells, and hyper-immunoglobulin E. Alteration in ARPC1B protein structure led to absent/low expression by flow cytometry and confocal microscopy. This molecular defect was associated with the inability of patient-derived T cells to extend an actin-rich lamellipodia upon T-cell receptor (TCR) stimulation and to assemble an immunological synapse. ARPC1B-deficient T cells additionally displayed impaired TCR-mediated proliferation and SDF1-α-directed migration. Gene transfer of ARPC1B in patients' T cells using a lentiviral vector restored both ARPC1B expression and T-cell proliferation in vitro. In 2 of the patients, in vivo somatic reversion restored ARPC1B expression in a fraction of lymphocytes and was associated with a skewed TCR repertoire. In 1 revertant patient, memory CD8+ T cells expressing normal levels of ARPC1B displayed improved T-cell migration. Inherited ARPC1B deficiency therefore alters T-cell cytoskeletal dynamics and functions, contributing to the clinical features of CID.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
ARPC1B is a key factor for the assembly and maintenance of the ARP2/3 complex that is involved in actin branching from an existing filament. Germline biallelic mutations in ARPC1B have been recently described in 6 patients with clinical features of combined immunodeficiency (CID), whose neutrophils and platelets but not T lymphocytes were studied. We hypothesized that ARPC1B deficiency may also lead to cytoskeleton and functional defects in T cells. We have identified biallelic mutations in ARPC1B in 6 unrelated patients with early onset disease characterized by severe infections, autoimmune manifestations, and thrombocytopenia. Immunological features included T-cell lymphopenia, low numbers of naïve T cells, and hyper-immunoglobulin E. Alteration in ARPC1B protein structure led to absent/low expression by flow cytometry and confocal microscopy. This molecular defect was associated with the inability of patient-derived T cells to extend an actin-rich lamellipodia upon T-cell receptor (TCR) stimulation and to assemble an immunological synapse. ARPC1B-deficient T cells additionally displayed impaired TCR-mediated proliferation and SDF1-α-directed migration. Gene transfer of ARPC1B in patients' T cells using a lentiviral vector restored both ARPC1B expression and T-cell proliferation in vitro. In 2 of the patients, in vivo somatic reversion restored ARPC1B expression in a fraction of lymphocytes and was associated with a skewed TCR repertoire. In 1 revertant patient, memory CD8+ T cells expressing normal levels of ARPC1B displayed improved T-cell migration. Inherited ARPC1B deficiency therefore alters T-cell cytoskeletal dynamics and functions, contributing to the clinical features of CID. |
2017
|
Choi, Seeyoung; Cornall, Richard; Lesourne, Renaud; Love, Paul E. THEMIS: Two Models, Different Thresholds Journal Article In: Trends Immunol, vol. 38, no. 9, pp. 622–632, 2017, ISSN: 1471-4981. @article{choi_themis_2017b,
title = {THEMIS: Two Models, Different Thresholds},
author = {Choi, Seeyoung and Cornall, Richard and Lesourne, Renaud and Love, Paul E.},
doi = {10.1016/j.it.2017.06.006},
issn = {1471-4981},
year = {2017},
date = {2017-01-01},
journal = {Trends Immunol},
volume = {38},
number = {9},
pages = {622--632},
abstract = {THEMIS, a recently identified T-lineage-restricted protein, is the founding member of a large metazoan protein family. Gene inactivation studies have revealed a critical requirement for THEMIS during thymocyte positive selection, implicating THEMIS in signaling downstream of the T cell antigen receptor (TCR), but the mechanistic underpinnings of THEMIS function have remained elusive. A previous model posited that THEMIS prevents thymocytes from inappropriately crossing the positive/negative selection threshold by dampening TCR signaling. However, new data suggest an alternative model where THEMIS enhances TCR signaling enabling thymocytes to reach the threshold for positive selection, avoiding death by neglect. We review the data supporting each model and conclude that the preponderance of evidence favors an enhancing function for THEMIS in TCR signaling.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
THEMIS, a recently identified T-lineage-restricted protein, is the founding member of a large metazoan protein family. Gene inactivation studies have revealed a critical requirement for THEMIS during thymocyte positive selection, implicating THEMIS in signaling downstream of the T cell antigen receptor (TCR), but the mechanistic underpinnings of THEMIS function have remained elusive. A previous model posited that THEMIS prevents thymocytes from inappropriately crossing the positive/negative selection threshold by dampening TCR signaling. However, new data suggest an alternative model where THEMIS enhances TCR signaling enabling thymocytes to reach the threshold for positive selection, avoiding death by neglect. We review the data supporting each model and conclude that the preponderance of evidence favors an enhancing function for THEMIS in TCR signaling. |
Cheng, D.; Deobagkar-Lele, M.; Zvezdova, E.; Choi, S.; Uehara, S.; Baup, D.; Bennett, S. C.; Bull, K. R.; Crockford, T. L.; Ferry, H.; Warzecha, C.; Marcellin, M.; de Peredo, A. G.; Lesourne, R.; Anzilotti, C.; Love, P. E.; Cornall, R. J. Themis2 lowers the threshold for B cell activation during positive selection Journal Article In: Nat Immunol, vol. 18, no. 2, pp. 205-213, 2017, ISSN: 1529-2916 (Electronic)
1529-2908 (Linking). @article{RN6b,
title = {Themis2 lowers the threshold for B cell activation during positive selection},
author = {Cheng, D. and Deobagkar-Lele, M. and Zvezdova, E. and Choi, S. and Uehara, S. and Baup, D. and Bennett, S. C. and Bull, K. R. and Crockford, T. L. and Ferry, H. and Warzecha, C. and Marcellin, M. and de Peredo, A. G. and Lesourne, R. and Anzilotti, C. and Love, P. E. and Cornall, R. J.},
url = {http://www.ncbi.nlm.nih.gov/pubmed/27992403},
doi = {10.1038/ni.3642},
issn = {1529-2916 (Electronic)
1529-2908 (Linking)},
year = {2017},
date = {2017-01-01},
journal = {Nat Immunol},
volume = {18},
number = {2},
pages = {205-213},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
|
Choi, S.; Warzecha, C.; Zvezdova, E.; Lee, J.; Argenty, J.; Lesourne, R.; Aravind, L.; Love, P. E. THEMIS enhances TCR signaling and enables positive selection by selective inhibition of the phosphatase SHP-1 Journal Article In: Nat Immunol, vol. 18, no. 4, pp. 433-441, 2017, ISSN: 1529-2916 (Electronic)
1529-2908 (Linking). @article{RN5b,
title = {THEMIS enhances TCR signaling and enables positive selection by selective inhibition of the phosphatase SHP-1},
author = {Choi, S. and Warzecha, C. and Zvezdova, E. and Lee, J. and Argenty, J. and Lesourne, R. and Aravind, L. and Love, P. E.},
url = {http://www.ncbi.nlm.nih.gov/pubmed/28250424},
doi = {10.1038/ni.3692},
issn = {1529-2916 (Electronic)
1529-2908 (Linking)},
year = {2017},
date = {2017-01-01},
journal = {Nat Immunol},
volume = {18},
number = {4},
pages = {433-441},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
|
Combedazou, A.; Choesmel-Cadamuro, V.; Gay, G.; Liu, J.; Dupre, L.; Ramel, D.; Wang, X. Myosin II governs collective cell migration behaviour downstream of guidance receptor signalling Journal Article In: J Cell Sci, vol. 130, no. 1, pp. 97-103, 2017, ISSN: 1477-9137 (Electronic)
0021-9533 (Linking). @article{RN10b,
title = {Myosin II governs collective cell migration behaviour downstream of guidance receptor signalling},
author = {Combedazou, A. and Choesmel-Cadamuro, V. and Gay, G. and Liu, J. and Dupre, L. and Ramel, D. and Wang, X.},
url = {http://www.ncbi.nlm.nih.gov/pubmed/27034137},
doi = {10.1242/jcs.179952},
issn = {1477-9137 (Electronic)
0021-9533 (Linking)},
year = {2017},
date = {2017-01-01},
journal = {J Cell Sci},
volume = {130},
number = {1},
pages = {97-103},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
|
Cotta-de-Almeida, V.; Dupre, L.; Savino, W. Editorial: T-Cell Migration in Health and Disease Journal Article In: Front Immunol, vol. 8, pp. 132, 2017, ISSN: 1664-3224 (Print)
1664-3224 (Linking). @article{RN11b,
title = {Editorial: T-Cell Migration in Health and Disease},
author = {Cotta-de-Almeida, V. and Dupre, L. and Savino, W.},
url = {http://www.ncbi.nlm.nih.gov/pubmed/28243238},
doi = {10.3389/fimmu.2017.00132},
issn = {1664-3224 (Print)
1664-3224 (Linking)},
year = {2017},
date = {2017-01-01},
journal = {Front Immunol},
volume = {8},
pages = {132},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
|
Garreau, A.; Blaize, G.; Argenty, J.; Rouquie, N.; Tourdes, A.; Wood, S. A.; Saoudi, A.; Lesourne, R. Grb2-Mediated Recruitment of USP9X to LAT Enhances Themis Stability following Thymic Selection Journal Article In: J Immunol, 2017, ISSN: 1550-6606 (Electronic)
0022-1767 (Linking). @article{RN1b,
title = {Grb2-Mediated Recruitment of USP9X to LAT Enhances Themis Stability following Thymic Selection},
author = {Garreau, A. and Blaize, G. and Argenty, J. and Rouquie, N. and Tourdes, A. and Wood, S. A. and Saoudi, A. and Lesourne, R.},
url = {http://www.ncbi.nlm.nih.gov/pubmed/28877990},
doi = {10.4049/jimmunol.1700566},
issn = {1550-6606 (Electronic)
0022-1767 (Linking)},
year = {2017},
date = {2017-01-01},
journal = {J Immunol},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
|
Pfajfer, L.; Seidel, M. G.; Houmadi, R.; Rey-Barroso, J.; Hirschmugl, T.; Salzer, E.; Anton, I. M.; Urban, C.; Schwinger, W.; Boztug, K.; Dupre, L. WIP deficiency severely affects human lymphocyte architecture during migration and synapse assembly Journal Article In: Blood, vol. 130, no. 17, pp. 1949-1953, 2017, ISSN: 1528-0020 (Electronic)
0006-4971 (Linking). @article{RN12b,
title = {WIP deficiency severely affects human lymphocyte architecture during migration and synapse assembly},
author = {Pfajfer, L. and Seidel, M. G. and Houmadi, R. and Rey-Barroso, J. and Hirschmugl, T. and Salzer, E. and Anton, I. M. and Urban, C. and Schwinger, W. and Boztug, K. and Dupre, L.},
url = {http://www.ncbi.nlm.nih.gov/pubmed/28903942},
doi = {10.1182/blood-2017-04-777383},
issn = {1528-0020 (Electronic)
0006-4971 (Linking)},
year = {2017},
date = {2017-01-01},
journal = {Blood},
volume = {130},
number = {17},
pages = {1949-1953},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
|
Duguet, F.; Locard-Paulet, M.; Marcellin, M.; Chaoui, K.; Bernard, I.; Andreoletti, O.; Lesourne, R.; Burlet-Schiltz, O.; Gonzalez de Peredo, A.; Saoudi, A. Proteomic Analysis of Regulatory T Cells Reveals the Importance of Themis1 in the Control of Their Suppressive Function Journal Article In: Mol Cell Proteomics, vol. 16, no. 8, pp. 1416-1432, 2017, ISSN: 1535-9484 (Electronic) 1535-9476 (Linking). @article{RN4b,
title = {Proteomic Analysis of Regulatory T Cells Reveals the Importance of Themis1 in the Control of Their Suppressive Function},
author = {Duguet, F. and Locard-Paulet, M. and Marcellin, M. and Chaoui, K. and Bernard, I. and Andreoletti, O. and Lesourne, R. and Burlet-Schiltz, O. and Gonzalez de Peredo, A. and Saoudi, A.},
url = {http://www.ncbi.nlm.nih.gov/pubmed/28373295},
doi = {10.1074/mcp.M116.062745},
issn = {1535-9484 (Electronic) 1535-9476 (Linking)},
year = {2017},
date = {2017-01-01},
journal = {Mol Cell Proteomics},
volume = {16},
number = {8},
pages = {1416-1432},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
|
2016
|
Lamsoul, Isabelle; Uttenweiler-Joseph, Sandrine; Moog-Lutz, Christel; Lutz, Pierre G. Cullin 5-RING E3 ubiquitin ligases, new therapeutic targets? Journal Article In: Biochimie, vol. 122, pp. 339–347, 2016, ISSN: 1638-6183. @article{lamsoul_cullin_2016,
title = {Cullin 5-RING E3 ubiquitin ligases, new therapeutic targets?},
author = {Lamsoul, Isabelle and Uttenweiler-Joseph, Sandrine and Moog-Lutz, Christel and Lutz, Pierre G.},
doi = {10.1016/j.biochi.2015.08.003},
issn = {1638-6183},
year = {2016},
date = {2016-03-01},
journal = {Biochimie},
volume = {122},
pages = {339--347},
abstract = {Ubiquitylation is a reversible post-translational modification of proteins that controls a myriad of functions and cellular processes. It occurs through the sequential action of three distinct enzymes. E3 ubiquitin ligases (E3s) play the role of conductors of the ubiquitylation pathway making them attractive therapeutic targets. This review is dedicated to the largest family of multimeric E3s, the Cullin-RING E3 (CRL) family and more specifically to cullin 5 based CRLs that remains poorly characterized.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
Ubiquitylation is a reversible post-translational modification of proteins that controls a myriad of functions and cellular processes. It occurs through the sequential action of three distinct enzymes. E3 ubiquitin ligases (E3s) play the role of conductors of the ubiquitylation pathway making them attractive therapeutic targets. This review is dedicated to the largest family of multimeric E3s, the Cullin-RING E3 (CRL) family and more specifically to cullin 5 based CRLs that remains poorly characterized. |
Salzer, E.; Cagdas, D.; Hons, M.; Mace, E. M.; Garncarz, W.; Petronczki, O. Y.; Platzer, R.; Pfajfer, L.; Bilic, I.; Ban, S. A.; Willmann, K. L.; Mukherjee, M.; Supper, V.; Hsu, H. T.; Banerjee, P. P.; Sinha, P.; McClanahan, F.; Zlabinger, G. J.; Pickl, W. F.; Gribben, J. G.; Stockinger, H.; Bennett, K. L.; Huppa, J. B.; Dupre, L.; Sanal, O.; Jager, U.; Sixt, M.; Tezcan, I.; Orange, J. S.; Boztug, K. RASGRP1 deficiency causes immunodeficiency with impaired cytoskeletal dynamics Journal Article In: Nat Immunol, vol. 17, no. 12, pp. 1352-1360, 2016, ISSN: 1529-2916 (Electronic)
1529-2908 (Linking). @article{RN14,
title = {RASGRP1 deficiency causes immunodeficiency with impaired cytoskeletal dynamics},
author = {Salzer, E. and Cagdas, D. and Hons, M. and Mace, E. M. and Garncarz, W. and Petronczki, O. Y. and Platzer, R. and Pfajfer, L. and Bilic, I. and Ban, S. A. and Willmann, K. L. and Mukherjee, M. and Supper, V. and Hsu, H. T. and Banerjee, P. P. and Sinha, P. and McClanahan, F. and Zlabinger, G. J. and Pickl, W. F. and Gribben, J. G. and Stockinger, H. and Bennett, K. L. and Huppa, J. B. and Dupre, L. and Sanal, O. and Jager, U. and Sixt, M. and Tezcan, I. and Orange, J. S. and Boztug, K.},
url = {http://www.ncbi.nlm.nih.gov/pubmed/27776107},
doi = {10.1038/ni.3575},
issn = {1529-2916 (Electronic)
1529-2908 (Linking)},
year = {2016},
date = {2016-01-01},
journal = {Nat Immunol},
volume = {17},
number = {12},
pages = {1352-1360},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
|
Zvezdova, E.; Mikolajczak, J.; Garreau, A.; Marcellin, M.; Rigal, L.; Lee, J.; Choi, S.; Blaize, G.; Argenty, J.; Familiades, J.; Li, L.; Gonzalez de Peredo, A.; Burlet-Schiltz, O.; Love, P. E.; Lesourne, R. Themis1 enhances T cell receptor signaling during thymocyte development by promoting Vav1 activity and Grb2 stability Journal Article In: Sci Signal, vol. 9, no. 428, pp. ra51, 2016, ISSN: 1937-9145 (Electronic)
1945-0877 (Linking). @article{RN7b,
title = {Themis1 enhances T cell receptor signaling during thymocyte development by promoting Vav1 activity and Grb2 stability},
author = {Zvezdova, E. and Mikolajczak, J. and Garreau, A. and Marcellin, M. and Rigal, L. and Lee, J. and Choi, S. and Blaize, G. and Argenty, J. and Familiades, J. and Li, L. and Gonzalez de Peredo, A. and Burlet-Schiltz, O. and Love, P. E. and Lesourne, R.},
url = {http://www.ncbi.nlm.nih.gov/pubmed/27188442},
doi = {10.1126/scisignal.aad1576},
issn = {1937-9145 (Electronic)
1945-0877 (Linking)},
year = {2016},
date = {2016-01-01},
journal = {Sci Signal},
volume = {9},
number = {428},
pages = {ra51},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
|
2015
|
Pedros, C.; Gaud, G.; Bernard, I.; Kassem, S.; Chabod, M.; Lagrange, D.; Andreoletti, O.; Dejean, A. S.; Lesourne, R.; Fournie, G. J.; Saoudi, A. An Epistatic Interaction between Themis1 and Vav1 Modulates Regulatory T Cell Function and Inflammatory Bowel Disease Development Journal Article In: J Immunol, vol. 195, no. 4, pp. 1608-16, 2015, ISSN: 1550-6606 (Electronic)
0022-1767 (Linking). @article{RN8,
title = {An Epistatic Interaction between Themis1 and Vav1 Modulates Regulatory T Cell Function and Inflammatory Bowel Disease Development},
author = {Pedros, C. and Gaud, G. and Bernard, I. and Kassem, S. and Chabod, M. and Lagrange, D. and Andreoletti, O. and Dejean, A. S. and Lesourne, R. and Fournie, G. J. and Saoudi, A.},
url = {http://www.ncbi.nlm.nih.gov/pubmed/26163585},
doi = {10.4049/jimmunol.1402562},
issn = {1550-6606 (Electronic)
0022-1767 (Linking)},
year = {2015},
date = {2015-01-01},
journal = {J Immunol},
volume = {195},
number = {4},
pages = {1608-16},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
|
Malet-Engra, G.; Yu, W.; Oldani, A.; Rey-Barroso, J.; Gov, N. S.; Scita, G.; Dupre, L. Collective cell motility promotes chemotactic prowess and resistance to chemorepulsion Journal Article In: Curr Biol, vol. 25, no. 2, pp. 242-50, 2015, ISSN: 1879-0445 (Electronic)
0960-9822 (Linking). @article{RN19b,
title = {Collective cell motility promotes chemotactic prowess and resistance to chemorepulsion},
author = {Malet-Engra, G. and Yu, W. and Oldani, A. and Rey-Barroso, J. and Gov, N. S. and Scita, G. and Dupre, L.},
url = {http://www.ncbi.nlm.nih.gov/pubmed/25578904},
doi = {10.1016/j.cub.2014.11.030},
issn = {1879-0445 (Electronic)
0960-9822 (Linking)},
year = {2015},
date = {2015-01-01},
journal = {Curr Biol},
volume = {25},
number = {2},
pages = {242-50},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
|
Dupre, L.; Houmadi, R.; Tang, C.; Rey-Barroso, J. T Lymphocyte Migration: An Action Movie Starring the Actin and Associated Actors Journal Article In: Front Immunol, vol. 6, pp. 586, 2015, ISSN: 1664-3224 (Print)
1664-3224 (Linking). @article{RN18b,
title = {T Lymphocyte Migration: An Action Movie Starring the Actin and Associated Actors},
author = {Dupre, L. and Houmadi, R. and Tang, C. and Rey-Barroso, J.},
url = {http://www.ncbi.nlm.nih.gov/pubmed/26635800},
doi = {10.3389/fimmu.2015.00586},
issn = {1664-3224 (Print)
1664-3224 (Linking)},
year = {2015},
date = {2015-01-01},
journal = {Front Immunol},
volume = {6},
pages = {586},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
|
Vasconcelos, Z.; Muller, S.; Guipouy, D.; Yu, W.; Christophe, C.; Gadat, S.; Valitutti, S.; Dupre, L. Individual Human Cytotoxic T Lymphocytes Exhibit Intraclonal Heterogeneity during Sustained Killing Journal Article In: Cell Rep, vol. 11, no. 9, pp. 1474-85, 2015, ISSN: 2211-1247 (Electronic). @article{RN20b,
title = {Individual Human Cytotoxic T Lymphocytes Exhibit Intraclonal Heterogeneity during Sustained Killing},
author = {Vasconcelos, Z. and Muller, S. and Guipouy, D. and Yu, W. and Christophe, C. and Gadat, S. and Valitutti, S. and Dupre, L.},
url = {http://www.ncbi.nlm.nih.gov/pubmed/26027932},
doi = {10.1016/j.celrep.2015.05.002},
issn = {2211-1247 (Electronic)},
year = {2015},
date = {2015-01-01},
journal = {Cell Rep},
volume = {11},
number = {9},
pages = {1474-85},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
|
Cotta-de-Almeida, V.; Dupre, L.; Guipouy, D.; Vasconcelos, Z. Signal Integration during T Lymphocyte Activation and Function: Lessons from the Wiskott-Aldrich Syndrome Journal Article In: Front Immunol, vol. 6, pp. 47, 2015, ISSN: 1664-3224 (Print)
1664-3224 (Linking). @article{RN17b,
title = {Signal Integration during T Lymphocyte Activation and Function: Lessons from the Wiskott-Aldrich Syndrome},
author = {Cotta-de-Almeida, V. and Dupre, L. and Guipouy, D. and Vasconcelos, Z.},
url = {http://www.ncbi.nlm.nih.gov/pubmed/25709608},
doi = {10.3389/fimmu.2015.00047},
issn = {1664-3224 (Print)
1664-3224 (Linking)},
year = {2015},
date = {2015-01-01},
journal = {Front Immunol},
volume = {6},
pages = {47},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
|
Christophe, C.; Muller, S.; Rodrigues, M.; Petit, A. E.; Cattiaux, P.; Dupre, L.; Gadat, S.; Valitutti, S. A biased competition theory of cytotoxic T lymphocyte interaction with tumor nodules Journal Article In: PLoS One, vol. 10, no. 3, pp. e0120053, 2015, ISSN: 1932-6203 (Electronic)
1932-6203 (Linking). @article{RN16b,
title = {A biased competition theory of cytotoxic T lymphocyte interaction with tumor nodules},
author = {Christophe, C. and Muller, S. and Rodrigues, M. and Petit, A. E. and Cattiaux, P. and Dupre, L. and Gadat, S. and Valitutti, S.},
url = {http://www.ncbi.nlm.nih.gov/pubmed/25815811},
doi = {10.1371/journal.pone.0120053},
issn = {1932-6203 (Electronic)
1932-6203 (Linking)},
year = {2015},
date = {2015-01-01},
journal = {PLoS One},
volume = {10},
number = {3},
pages = {e0120053},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
|
Spinner, Camille A.; Uttenweiler-Joseph, Sandrine; Metais, Arnaud; Stella, Alexandre; Burlet-Schiltz, Odile; Moog-Lutz, Christel; Lamsoul, Isabelle; Lutz, Pierre G. Substrates of the ASB2α E3 ubiquitin ligase in dendritic cells Journal Article In: Scientific Reports, vol. 5, no. 1, pp. 16269, 2015, ISSN: 2045-2322, (Number: 1
Publisher: Nature Publishing Group). @article{spinner_substrates_2015,
title = {Substrates of the ASB2α E3 ubiquitin ligase in dendritic cells},
author = {Spinner, Camille A. and Uttenweiler-Joseph, Sandrine and Metais, Arnaud and Stella, Alexandre and Burlet-Schiltz, Odile and Moog-Lutz, Christel and Lamsoul, Isabelle and Lutz, Pierre G.},
url = {http://www.nature.com/articles/srep16269},
doi = {10.1038/srep16269},
issn = {2045-2322},
year = {2015},
date = {2015-01-01},
urldate = {2020-09-30},
journal = {Scientific Reports},
volume = {5},
number = {1},
pages = {16269},
abstract = {Conventional dendritic cells (cDCs) comprise distinct populations with specialized immune functions that are mediators of innate and adaptive immune responses. Transcriptomic and proteomic approaches have been used so far to identify transcripts and proteins that are differentially expressed in these subsets to understand the respective functions of cDCs subsets. Here, we showed that the Cullin 5-RING E3 ubiquitin ligase (E3) ASB2α, by driving degradation of filamin A (FLNa) and filamin B (FLNb), is responsible for the difference in FLNa and FLNb abundance in the different spleen cDC subsets. Importantly, the ability of these cDC subsets to migrate correlates with the level of FLNa. Furthermore, our results strongly point to CD4 positive and double negative cDCs as distinct populations. Finally, we develop quantitative global proteomic approaches to identify ASB2α substrates in DCs using ASB2 conditional knockout mice. As component of the ubiquitin-proteasome system (UPS) are amenable to pharmacological manipulation, these approaches aimed to the identification of E3 substrates in physiological relevant settings could potentially lead to novel targets for therapeutic strategies.},
note = {Number: 1
Publisher: Nature Publishing Group},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
Conventional dendritic cells (cDCs) comprise distinct populations with specialized immune functions that are mediators of innate and adaptive immune responses. Transcriptomic and proteomic approaches have been used so far to identify transcripts and proteins that are differentially expressed in these subsets to understand the respective functions of cDCs subsets. Here, we showed that the Cullin 5-RING E3 ubiquitin ligase (E3) ASB2α, by driving degradation of filamin A (FLNa) and filamin B (FLNb), is responsible for the difference in FLNa and FLNb abundance in the different spleen cDC subsets. Importantly, the ability of these cDC subsets to migrate correlates with the level of FLNa. Furthermore, our results strongly point to CD4 positive and double negative cDCs as distinct populations. Finally, we develop quantitative global proteomic approaches to identify ASB2α substrates in DCs using ASB2 conditional knockout mice. As component of the ubiquitin-proteasome system (UPS) are amenable to pharmacological manipulation, these approaches aimed to the identification of E3 substrates in physiological relevant settings could potentially lead to novel targets for therapeutic strategies. |
Amin, R.; Mourcin, F.; Uhel, F.; Pangault, C.; Ruminy, P.; Dupre, L.; Guirriec, M.; Marchand, T.; Fest, T.; Lamy, T.; Tarte, K. DC-SIGN-expressing macrophages trigger activation of mannosylated IgM B-cell receptor in follicular lymphoma Journal Article In: Blood, vol. 126, no. 16, pp. 1911-20, 2015, ISSN: 1528-0020 (Electronic)
0006-4971 (Linking). @article{RN15b,
title = {DC-SIGN-expressing macrophages trigger activation of mannosylated IgM B-cell receptor in follicular lymphoma},
author = {Amin, R. and Mourcin, F. and Uhel, F. and Pangault, C. and Ruminy, P. and Dupre, L. and Guirriec, M. and Marchand, T. and Fest, T. and Lamy, T. and Tarte, K.},
url = {http://www.ncbi.nlm.nih.gov/pubmed/26272216},
doi = {10.1182/blood-2015-04-640912},
issn = {1528-0020 (Electronic)
0006-4971 (Linking)},
year = {2015},
date = {2015-01-01},
journal = {Blood},
volume = {126},
number = {16},
pages = {1911-20},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
|